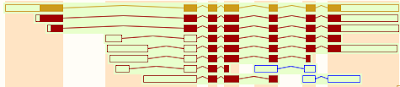
The transcripts of some genes can be alternatively spliced to produce more than one biologically functional product (e.g. proteins). There are several well-documented examples in the scientific literature but they are not common. There are probably fewer than 500 human genes (2.5%) that exhibit true alternative splicing where the alternate gene product has been conclusively shown to exist and be biologically functional.
However, it's easy to detect multiple examples of unusually spliced transcripts of humans genes. The vast majority of these splice variants are present at less than one copy per cell, are rapidly degraded, and not conserved in closely related species. That has led to the idea that they are simply the result of splicing errors, a conclusion that's reinforced by solid evidence that splicing is error prone.
It's unfortunate that all these splice variants are assumed to be real examples of alternative splicing leading to the widely held view that more than 90% of human protein-coding genes are alternatively spliced. This false claim is used as a way of getting around the Deflated Ego Problem by assuming that the "shockingly" small number of genes in humans is explained by the fact that humans have evolved mechanisms for producing up to one hundred thousand distinct proteins from only 20,000 protein-coding genes.In recent years, many scientists have come to realize that the role of alternative splicing has been greatly exaggerated. If you're interested in learning more, I cover the controversy on pages 154-169 in my book and in numerous blog posts (see below).
Unfortunately, there's a Nature editor who didn't get the message so they perpetuated the standard misinformation in a recent (May 21, 2025) editorial in Nature Reviews Genetics.
Anonymous (2025) RNA splicing — a central layer of gene regulation. Nat Rev Genet 26:369–370 [doi: 10.1038/s41576-025-00846-x]
... Splicing is essential for the accurate translation of DNA sequence information and comes with the added perk of generating transcriptomic and proteomic diversity in the form of alternative splicing — that is, the regulated inclusion or exclusion of exons. Alternative splicing greatly expands the coding potential of the genome; more than 95% of human multi-intron genes undergo alternative splicing, producing mRNA isoforms that can differ in coding sequence, regulatory elements or untranslated regions. These isoforms can influence mRNA stability, localization and translation output, thereby modulating cellular function.
... The ability of a single gene to produce several, functionally distinct protein isoforms through alternative splicing could enable organisms to rapidly adapt to changing environments. By enabling the sequencing of full-length transcripts, long-read sequencing data have yielded a more complete picture of alternative splicing. Subsequent comparative transcriptomic studies have revealed striking differences in the extent of alternative splicing between eukaryotes. Indeed, recent studies suggest that heritable variation in patterns of alternative splicing contributes to adaptive evolutionary chang.
Wouldn't it be nice if some leading researchers in the field wrote a scathing letter to Nature about the propagation of such misinformation? Does anyone know who to contact at Nature if you want to register a complaint?
Blog posts on alternative splicing
- The number of splice variants in a species correlates inversely with the population size - what does that mean?
- Splicing errors or alternative splicing?
- Alternative splicing and evolution
- Alternative splicing: function vs noise
- The frequency of splicing errors reflects the balance between selection and drift
- Alternative splicing in the nematode C. elegans
- The persistent myth of alternative splicing
- The textbook view of alternative splicing
- Are splice variants functional or noise?
- Debating alternative splicing (part I)
- Debating alternative splicing (part II)
- Debating alternative splicing (Part III)
- Debating alternative splicing (Part IV)