It's going to be extremely difficult, perhaps impossible, to merge the new complete human genome sequence with the current standard reference genome.
The source DNA for the new telomere-to-telomere (T2T) human genome sequence was a cell line derived from a molar pregnancy. This meant that the DNA was essentially haploid, thus avoiding the complications of sequencing diploid DNA which contains two highly similar but different genomes. The cell line, CHM13, lacks a Y chromosome but that's trivial since a complete T2T sequence of a Y chromosome will soon be published and it can be added to the T2T-CHM13 genome sequence [Telomere-to-telomere sequencing of a complete human genome].
The current standard reference genome is CHCh38.p13 (Feb. 28, 2019). It is a vitual sequence derived from a number of individuals living near Buffalo, New York. Since publication of the first human genome sequences, there have been thousands of others and none of them match the standard reference genome because of polymorphic SNPs and various deletions and insertions (indels). Some of these deletions and insertions can be very large (e.g. segmental duplications) so that no two human genome sequences are identical or even the same size (other than identical twins).
None of this is new. We all know that the standard reference genome is just that, a reference genome. We all know that there's a huge amount of variation between individuals; it's exactly what you expect for a dynamic genome where most changes, including deletions and insertions, are not restrained by purifying selection. It's good evidence that most of our genome is junk.
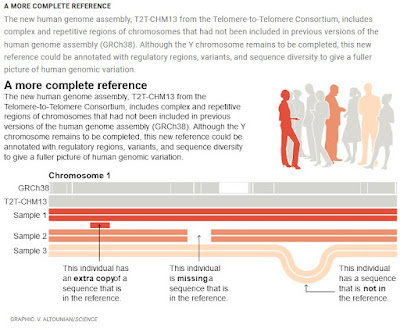
A lot of this variation at the level of SNPs and short indels can be handled by annotating the standard reference genome. Larger insertions and deletions, and chromosomal rearrangements, require a supplemental database that can be linked to the standard reference genome. This is one way to deal with the "pangenome"—the complete sequences of every known genome.
But this isn't as easy as it seems and it's especially complicated with the new complete sequence. A good discussion of the problem with integrating the T2T-CHM13 assembly can be found in a short essay by Deanna Church in the same issue of Science that contains the new sequence papers (Church, 2022). The new assembly corrects some errors in the CHCh38 assembly and adds an extra 8% of the genome. What that means is that the extensive annotation in the standard reference genome, can't be easily transferred to the T2T-CHM13 assembly because, for one thing, the numbering of the bases is very different. In addition, there are extra sequences in T2T-CHM13 that have to annotated. If they are duplications then you have to figure out which copy corresponds to the CHCh38 version and that's going to take time. Church shows some of the issues in a figure.
Keep in mind that the sequence of the standard reference genome is important but the annotation is equally important. A lot of genomics work relies on accurate annotation of coding regions, regulatory sequences, transposons, origins of replication, and a host of other markers. The reference genome is routinely scanned to extract this information. Genome wide association studies (GWAS) rely on the annotation. This annotation is the product of 22 years of work by hundreds of scientists and it's not going to be easy to extend it to the T2T-CHM13 genome.
- What do we do with two different human genome reference sequences?
- Karen Miga and the telomere-to-telomere consortium
- A complete human genome sequence (2022)
- Epigenetic markers in the last 8% of the human genome sequence
- Segmental duplications in the human genome
- Transcription activity in repeat regions of the human genome
- Genetic variation and the complete human genome sequence
Church, D.M. (2022) A next-generation human genome sequence. Science 376:34-35. [doi: 10.1126/science.abo5367]
No comments :
Post a Comment