Most of the genes in eukaryotes contain introns that are removed by splicing during processing of the primary transcript. In some cases the gene produces two different functional RNAs due to differential splicing of the introns. If the product is mRNA then two different versions of the protein can be made as shown in the figure from my book What's in Your Genome? This mechanism is known as alternative splicing.
True alternative splicing is rare—less than 5% of all genes are alternatively spliced.1 However, when you analyze all of the transcripts in a tissue you will invariably detect many transcripts from junk DNA and many low abundance splice variants. Those transcripts and splice variants are due to transcription errors and splicing errors. Splicing errors arise from the presence of weak splice sites that are occasionally recognized by the normal spliceosome or by the splice factors responsible for true alternative splicing.
Laurent Duret and his colleagues wondered why those weak spurious splice sites aren't purged from the genome by negative selection and they postulated that natural selection isn't powerful enough to purge the genome of these sites in species with small population sizes. Conversely, in species with large population sizes, many of them will be removed by negative selection so there should be fewer spurious splice variants in species with large population sizes.
The results are published in a revised manuscript posted to the bioRxiv site a few weeks ago (Sept. 26, 2023).2 Normally I would wait until the paper appeared in a peer-reviewed journal but this time I want to make a few comments before it gets published.
Bénitière, F., Necsulea, A. and Duret, L. (2023) Random genetic drift sets an upper limit on mRNA splicing accuracy in metazoans. bioRxiv:2022.2012. 2009.519597. [doi: 10.1101/2022.12.09.519597]
Most eukaryotic genes undergo alternative splicing (AS), but the overall functional significance of this process remains a controversial issue. It has been noticed that the complexity of organisms (assayed by the number of distinct cell types) correlates positively with their genome-wide AS rate. This has been interpreted as evidence that AS plays an important role in adaptive evolution by increasing the functional repertoires of genomes. However, this observation also fits with a totally opposite interpretation: given that ‘complex’ organisms tend to have small effective population sizes (N e), they are expected to be more affected by genetic drift, and hence more prone to accumulate deleterious mutations that decrease splicing accuracy. Thus, according to this “drift barrier” theory, the elevated AS rate in complex organisms might simply result from a higher splicing error rate. To test this hypothesis, we analyzed 3,496 transcriptome sequencing samples to quantify AS in 53 metazoan species spanning a wide range of N e values. Our results show a negative correlation between N e proxies and the genome-wide AS rates among species, consistent with the drift barrier hypothesis. This pattern is dominated by low abundance isoforms, which represent the vast majority of the splice variant repertoire. We show that these low abundance isoforms are depleted in functional AS events, and most likely correspond to errors. Conversely, the AS rate of abundant isoforms, which are relatively enriched in functional AS events, tends to be lower in more complex species. All these observations are consistent with the hypothesis that variation in AS rates across metazoans reflects the limits set by drift on the capacity of selection to prevent gene expression errors.
The paper is a bit complicated so I'll simplify a great deal. You'll have to read it yourself to get all the details.
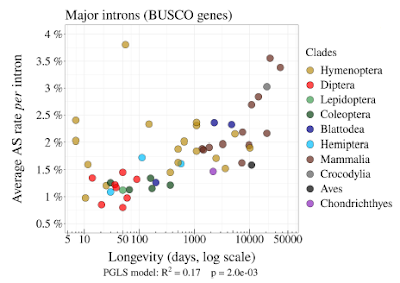
They looked at the number of splice variants per intron in a set of 978 orthologous genes in vertebrates and insects (BUSCO genes). Then they used a number of proxies for population size including the average longevity of individuals in the species. (This is more accurate than you might think.) When you plot the number of variants vs longevity you get the following graph.
They interpret this to mean that their hypothesis is confirmed; species with small population sizes (greater longevity) have more splice variants than species with large population sizes and this is "consistent with the hypothesis that variation in AS rates across metazoans reflects the limits set by drift on the capacity of selection to prevent gene expression errors."
There are all kinds of issues with this type of study but the authors seem to have addressed most of them. Note that the graph plots the number of splice variants per intron but this can be a bit deceptive since the number of introns per gene differs significantly from only 2.8 introns per gene in Diptera to an average of 8.4 introns per gene in vertebrates. The correlation between splice variants per gene and population size (longevity) is even more striking whether you use just the BUSCO genes or all protein coding genes.
The authors assume that almost all of these splice variants are due to processing errors where aberrant splicing is caused by the recognition of incorrect splice sites. Thus, "... these observations fit very well with a model where variation in AS rate across species is entirely driven by variation in the efficacy of selection against splicing errors." The idea is that spurious splice sites are removed more often by natural selection in species with large population sizes (e.g. insects) than in species with small population sizes (e.g. mammals)
I think there's a better explanation. Genome size also correlates with population size and the larger the genome the more junk DNA. Introns are mostly junk so large genomes have more introns and larger introns. Thus, in mammals there are lots of introns and they can be huge. This means there's a lot more opportunity for aberrant splicing at spurious splice sites. In species with smaller genomes, such as Diptera, here are fewer introns and they are much smaller than the mammalian introns. The target size for spurious splice sites is much smaller so there are fewer splicing errors per intron and a lot fewer per gene.
I looked up the genome sizes for these species on Ryan Gregory's Animal Genome Size Database and the correlation between genome size and the number of splice variants seems to hold fairly well. There's some scatter but not enough to rule out a strong correlation between genome size and the amount of intron DNA in a gene. I attribute the huge number of splice variants in mammals to the presence of a large genome full of junk DNA, much of which is in introns. In other species with smaller genomes there will be fewer splicing errors because the introns are smaller.
There's a correlation between splicing errors and population size but it's not, in my opinion, due to direct selection against the sequence of aberrant splice sites in species with large populations as proposed by Bénitière et al., but by reduction in the size of introns and the number of extraneous weak splice sites. Thus, in my view, increases in population size could lead to selection against junk DNA and a reduction in genome size and, as a consequence (byproduct), there will be fewer transcription errors and fewer splicing errors.
1. This is not the place to argue this point. It's covered in my book and in many blog posts [Splicing errors or alternative splicing?].
2. The first version was posted in December, 2022.


4 comments :
Why is there selection against junk DNA though? Surely part of it is selection against maladaptive consequences of excess junk… such as an increase in splicing errors. I’m not sure how these two explanations are supposed to conflict with each other.
Is this correct?
"species with small population sizes (greater longevity) have more splice variants than species with smaller population sizes and this "
Great post, Larry!
@João: No the statement was not correct. I fixed it. Thanks.
To be fair they attribute this increase in apparent alternative splicing to an increase in the "rate of splicing errors" - these could be caused by weaker splice sites (and other motifs that direct splicing), excess intronic sequence or differences in the spliceosome. To be fair, it is likely all three. They could test the weaker splice site theory and look at splice site motif heterogeneity.
Post a Comment