The authors of a recent paper think we need a new term "spam DNA" to describe some features of the human genome.
Fagundes, N.J., Bisso-Machado, R., Figueiredo, P.I., Varal, M. and Zani, A.L. (2022) What We Talk About When We Talk About “Junk DNA”. Genome Biology and Evolution 14:evac055. [doi: 10.1093/gbe/evac055]
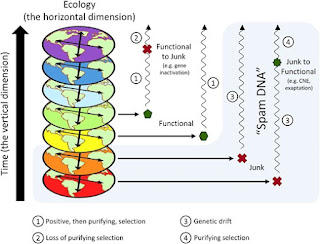
“Junk DNA” is a popular yet controversial concept that states that organisms carry in their genomes DNA that has no positive impact on their fitness. Nonetheless, biochemical functions have been identified for an increasing fraction of DNA elements traditionally seen as “Junk DNA”. These findings have been interpreted as fundamentally undermining the “Junk DNA” concept. Here, we reinforce previous arguments that this interpretation relies on an inadequate concept of biological function that does not consider the selected effect of a given genomic structure, which is central to the “Junk DNA” concept. Next, we suggest that another (though ignored) confounding factor is that the discussion about biological functions includes two different dimensions: a horizontal, ecological dimension that reflects how a given genomic element affects fitness in a specific time, and a vertical, temporal dimension that reflects how a given genomic element persisted along time. We suggest that “Junk DNA” should be used exclusively relative to the horizontal dimension, while for the vertical dimension, we propose a new term, “Spam DNA”, that reflects the fact that a given genomic element may persist in the genome even if not selected for on their origin. Importantly, these concepts are complementary. An element can be both “Spam DNA” and “Junk DNA”, and “Spam DNA” can also be recruited to perform evolved biological functions, as illustrated in processes of exaptation or constructive neutral evolution.
The authors are scientists at the Federal Univesity of Rio Grande do Sul in Brazil. They are concerened about the origins of junk DNA and whether true selected effect functions (strong selected effect = SSE) conflict with the definition of junk DNA. Here's how they put it,
Paradoxically as it may seem, under the SSE definition, elements that contribute positively to fitness and are maintained by purifying selection would still count as “junk” only because they did not originate as an adaptation.
This is essentially correct according to how many philosophers define selected effect functions but that issue was resolved by focusing on purifying selection as the important criterion and ignoring the history of the trait (= maintenance function, MF). There is only a 'paradox' if you stick to the philosophy definition of function (i.e. SSE) and even then, the paradox only exists if the SSE definition is the only way to identify junk DNA. [see: The Function Wars Part IX: Stefan Linquist on Causal Role vs Selected Effect] The authors recognize this since they include a good discussion of this other definition (MF) and its advantages. Nevertheless, they propose a new term called "spam DNA" to help clarify the problem.
"Spam DNA" represents every genomic element which has not been selected for during its origin in the genome, even if it currently participates in relevant biological functions.
All of the DNA in the light blue box is spam DNA. Note that it includes DNA that is currently functional as long as it originated from junk DNA as they define it. Also, some junk DNA isn't spam DNA as long as it arose from the inactivation of DNA that used to have a function. Thus, pseudogenes aren't junk and neither are bits and pieces of transposons.
This isn't helpful. The current debate is about how much of our genome is junk so who cares about the history of individual sequences? A significant amount of what we currently define as junk DNA may have come from once-active transposons but we may never be able to trace the history of each piece of junk DNA. Does it fall into the first category in the figure (functional to junk) or is it spam DNA? Is this really important? No,it is not.
Function Wars(My personal view of the meaning of function is described at the end of Part V.)
- On the Meaning of the Word "Function"
- The Function Wars: Part I
- The Function Wars: Part II
- The Function Wars: Part III
- The Function Wars: Part IV
- Restarting the function wars (The Function Wars Part V)
- The Function Wars Part VI: The problem with selected effect function
- The Function Wars Part VII: Function monism vs function pluralism
- The Function Wars Part VIII: Selected effect function and de novo genes
- The Function Wars Part IX: Stefan Linquist on Causal Role vs Selected Effect
- The Function Wars Part X: "Spam DNA"?

16 comments :
I discovered this blog very recently, 2 weeks, and I haven't read the full 'Function Wars' series thoroughly (I did skim through it to try and avoid missing obvious answers).
What I wonder is, do we (try to) consider possible beneficial effects of the unused/junk/spam DNA at population and evolutionary time scales? (I'm thinking in terms of its effects on the genome organization.) What prompted this line of questioning to me was that most vertebrates (if I remember correctly) have amounts of junk DNA with similar order of magnitude as humans, but some exotic branches (some fish maybe?) don't, and have a genome that's only a few percent the size of their brethren and their phylum. These rare branches would have lost the junk stuff over evolutionary time scales.
(I realize I'm not precise at all about the facts I'm mentioning, I'm a student and struggling to put it all together.)
So I'm thinking, if in most of the animal tree of life, presumably plants, fungi and others eukaryotes ditto, a large genome comprising much junk persisted, while we know it's possible to get rid of it, then it could be beneficial at some large population and time scales? I'm thinking of exon shuffling facilitation as a simple possible mechanism (all the dead transposons are great landing sites for the ones still alive), maybe other mechanisms I don't know of or that we haven't discovered yet.
My apologies if I'm miscontruing these questions as relevant to the debate at hand or missed that they are covered already..
If you want to learn something about comparatice genome sizes, I suggest the genome size database. That should bring a little rigor to your speculation.
What do you mean by "it's possible to get rid of it"?
What do you mean by "it's possible to get rid of it"?
Alas, what a lot of people mean by that is "if we get rid of it, the animal (or whatever) seems still to be alive". This implicitly recognizes only two values of fitness, 0 and 1. Once we recognize that the creature could look fine to us but have much lower fitness than the same species in nature, then we know we have to ask the question differently.
So if I understand this correctly, the authors propose to relabel all Junk DNA as Spam DNA, unless the sequence was instantly adaptive when it arose and later rendered inactive (which my gut feeling tells me may be uncommon)?
From skimming the paper, I understand that the authors believe this to be remedial against panadaptationist thinking, as it emphasizes that many current functional elements originate from previously neutral sequences. While that is a noble cause, I agree that it most likely will only add to the confusion, rather than clarify matters.
These posts made me think about all the junk in my garage that I keep just in case it is someday useful. Instead of going to the store I already have what I need. Does junk DNA help species evolve more quickly as the environment changes? It seems like it might take fewer mutations to make a pseudogene functional than to achieve the same functionality by another mechanism.
I also wonder about the sharp rhetorical distinction between junk and not junk. Presumably there are genes that are not completely useless but not useful enough that the selective pressure overwhelms the noise. They are on the road from gene to pseudogene. How long does that take? If the environment changes can they become genes again?
I'm not suggesting any teleology. In truth, my junk drawer is more a matter of me being too lazy to clean and organize than any real foresight. The usefulness is just an excuse I tell my spouse to avoid cleaning it out. And even when I think there's something I can use in all the junk, it's often just easier to go the store and buy a new one.
Does this analogy make any sense? Is there any evidence for junk, or on-the-road-to-junk becoming functional again?
The difficulty with any appeal to future use is that there is no current selection maintaining the trait. If selection is rare enough in time compared to individual generations, the evolution of that trait is dominated by the common situation. The K/T extinction selected for small size, but it exerted no selection until that moment, and large animals quickly re-appeared afterwards. So if some bit of junk occasionally becomes advantageous, this exerts no selection for the maintenance of large amounts of junk DNA. At most is might have a group selection effect, in which species with lots of junk would tend to go extinct less often than those with little junk, thus biasing the biota in favor of lots of junk. Group selection, however, is weak compared to within-population selection.
I believe a better analogy would be a ravine out back where you and the neighbors all dump trash, including good stuff you just don't need right now, from appliances to toothpicks. The metals rust. The organic objects, including paper, decompose. Animals scavenge, rearranging the junk and adding their own deposits. Weeds and decomposers grow and reproduce.
If you want something a neighbor threw out and you get there quickly, before it decomposes, you've got something new! This could be analogous to duplicated genes where both copies prove useful.
Is it possible that something useful will form spontaneously from all that junk? Yes. We do see occasional genes that seem to originate from a promoter that forms in front of a useful junk sequence. Is that likely? No. Not likely at all. Really, really not likely. But possible.
Getting rid of junk DNA isn't straightforward. There is no mechanism to read through DNA and say, "We need this, but we can throw this out." I don't know what happened with the fish, but the plant famous for its minimal DNA content and its relatives there have suffered massive DNA rearrangements. Those tend to leave most of the offspring dead (because essential sequences were lost) or unable to reproduce. Utricularia gibba and some of its relatives survived and U. gibba at least has a wonderfully efficient genome. But there is no straightforward way to get there from here.
My next post will be on Ford Doolittle's take on evolvability. It's possible to have selection for excess DNA as preparation for future evolution but that selection has to be at the level of species, not individuals.
The issue isn't whether we can speculate about such uses for junk DNA. It's whether there's any evidence to justify such speculation. As far as I know, de novo genes from junk are quite rare - about one every million years. I don't know of any evidence that these new genes conferred a selective advantage on a whole clade that allowed them to out compete other species in the same environment and that's what species sorting requires.
I think that rate of de novo coding gene generation (one every million years) might even be a bit generous. Particularly if we are talking about primates. If there is one de novo gene every million years, we might expect 4 or 5 de novo human genes not found in chimpanzee. Yet, despite many papers on the subject, we still haven't found one (which is not to say we won't find one eventually).
@Michael Tress
I was trying to err on the side of maximizing the rate. I was thinking of adding up all the possible candidates in the human genome and I couldn't come up with more than five total de novo genes that have a real possible function. They are all noncoding genes. I agree that there are no protein-coding genes.
One de novo gene per million years still could be a good general rate if we don't focus solely on the human lineage. Most organisms have much shorter generations than we do.
It is OK to use it as a ballpark number, but I imagine that de novo coding gene generation is going to vary a huge amount depending on the species. It is going to be much, much faster than one every million years in unicellular organisms (as an extreme example). But my point was more about the current absence of de novo human coding genes.
With some delay, I'd like to clarify:
- by getting rid of seemingly unused DNA, I don't mean through engineering, but that we can observe nature having done so, in some species;
- checking my source about the fish, that's actually Fugu, and my number was way off, Fugu having 40% as much DNA as its cousins, so not just "a few percent". The source that I had in mind being Albert's MBOC (page 222 in 6ed, "The Size of a Vertebrate Genome Reflects the Relative Rates of
DNA Addition and DNA Loss in a Lineage")
Hi Larry. Just found out these posts on "The Function Wars". Excellent reading and thanks for your thoughts on our piece.
Just a correction (perhaps it was not clear from the picture), for us, pseudogenes and active bits of transposons ARE junk DNA, as long as they are not maintained by purifying selection and do not contribute to organismal fitness. However, they are spam DNA if their persistence in the genome didn’t depend on adaptive processes. I would say that transposons are almost certainly spam DNA, and that pseudogenes are also spam DNA if their action was perfectly redundant with other genes before they were inactivated.
Perhaps an amendment we could make to Figure 1 is emphasizing that once junk DNA becomes functional (case four in Figure 1), there is also positive then purifying selection, not only purifying selection). In the end, I think I agree more with Brunet et al 2021 (https://doi.org/10.1016/j.shpsa.2021.03.005) than the text suggests. I see why they are somewhat reluctant in not mentioning positive selection as a fundamental process in originating function.
We just think it would be important to have different terms for accounting for the origin of an element in the genome and whether or not it contributes to fitness.
Thanks for the discussion
Cheers
"I see why they are somewhat reluctant in not mentioning positive selection as a fundamental process in originating function."
Please read:
I see why they are somewhat reluctant in not considering positive selection as a fundamental process in originating function.
Post a Comment