The term "function wars" refers to debates over the meaning of the word "function" in biology. It refers specifically to the discussion about junk DNA because junk DNA is defined as DNA that does not have a biological function. The wars were (re-)started when the ENCODE Consortium decided to use a stupid definition of function in order to prove that most of our genome was functional. This prompted a number of papers attempting to create a more meaningful definition.
None of them succeeded, in my opinion, because biology is messy and doesn't lend itself to precise definitions. Look how difficult it is to define a "gene," for example. Or "evolution."Nevertheless, some progress was made. Dan Graur has recently posted a summary of the two most important definitions of function [What does “function” mean in the context of evolution & what absurd situations may arise by using the wrong definition?]. The two definitions are "selected-effect" and "causal-role" (there are synonyms).
Selected-effect Function
At the risk of oversimplifying, a function is only a true biological function if it is selected. If you're trying to decide whether a particular DNA sequence has a biological function then you look to see if it is conserved. Sequence conservation is the primary indicator of function.
Causal-role Function
The other way of looking at function is to ask whether the DNA sequence does something. If it encodes a functional protein, for example, it has an obvious causal role. However, in this case it may also be conserved so an example like that doesn't distinguish between the two definitions. In fact, as Dan Graur (and others) point out, everything that meets the selected-effect definition (i.e. conserved) should also have a causal role.
The world is not inhabited exclusively by fools and when a subject arouses intense interest and debate, as this one has, something other than semantics is usually at stake.
Stephan Jay Gould (1982)We want to know about cases where a given DNA sequence does something (causal-role) but isn't conserved. Are those still examples of biological functions that don't meet the strict criterion of being conserved (i.e. selected by natural selection)? Conversely, are there examples of conserved sequences that don't have a real biological function?
The answer to both question is "yes" and that's why the selected-effect definition is not the definitive answer.
Before continuing, let me make it clear that sequence conservation is far and away the best indication of function that we have. It works almost all the time. As a first approximation, it's okay to insist on conservation as a crude definition of function as long as you aren't dogmatic about it. The selected-effect definition might be as good as it gets even though it's not perfect.
A sequence does something but isn't conserved
The best examples here are the ones mistakenly used by ENCODE. They identified transcription factor binding sites throughout the genome and suggested they all have a role in regulating gene expression. The second part of that claim is unproven—they may or may not have a biological role in regulation. It's very unlikely that they all have such a role.
The first part of the claim—that there are many transcription factor binding sites—is true. The DNA sites have a causal role in binding transcription factors. However, they are not conserved. When you look at the same locus in related species you often find that the sequence is different and the transcription factor will not bind to that site. Thus, the sequence might be functional according to the causal-role definition but it is not functional according to the selected-effect definition. In this case, the selected-effect definition trumps the causal-role definition and the DNA sequence does not have a biological function.1
Let's look at a more complicated example. Intron sequences are mostly junk but there's a minimal size of intron that's necessary for proper splicing. In most eukaryotes, it seems to be about 50 bp,4 or enough to form the loop of RNA that's required to bring the 5′ and 3′ splice sites together in the spliceosome. Most of that sequence isn't conserved but it definitely has a role to play in proper splicing.
Similarly, the spacing of transcription factor binding sites is also important in formation of loops of DNA that bring together the bound transcription factor and the RNA polymerase complex poised at the promoter. The spacer sequences are necessary but they aren't conserved.
Now, you could ague that the presence of a minimal spacer sequence IS conserved even though the actual DNA sequence is not. That's true but it seems to be stretching the selected-effect definition.
More importantly, there are many "bulk DNA" hypotheses that attempt to explain the presence of large amounts of superfluous DNA. For example, several workers have postulated that bulking up the genome leads to larger nuclei and larger cells. The bulk DNA has a function or sorts but the actual sequence is irrelevant.
These hypotheses may or may not be correct—I think they're wrong—but that's not the point. The point is that it's wrong to eliminate them by fiat on the grounds they don't meet the selected-effect definition of function. The arguments for and against bulk DNA hypotheses will have to be considered on their own merit regardless of any restriction imposed by blind allegiance to a specific definition of function.
There's a much more difficult example under this category. Active transposons can jump around in the genome. There are several examples of recent insertions in the human genome. Our closest relatives (chimps and bonobos) don't have a transposon at the orthologous locus.
The active transposon often carries genes for reverse transcriptase and some form of recombination enzyme for insertion and excision. The genes are transcribed and functional proteins are produced. This meets all the criteria for a causal-role definition of function but do these transposons really have a biological function or are they junk?
I maintain they are not junk, they are part of the functional DNA fraction of the genome. Other workers aren't so sure. They have developed a two-fold definition of biological function that distinguishes between function at the organism level and function at some other level. In this case, active transposon sequences don't meet the selected-effect definition at the level of the organism so they don't count as functional at that level.2 They may count as function at some other level, such as the intragenomic level or the level of selfish DNA, but that's not the same as the organismal level (Elliot et al., 2014; Doolittle et al., 2014).
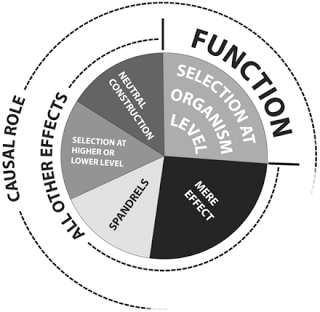
Doolittle et al. published a figure that illustrates their description of function. Look at the quadrant on the upper right—the one labelled "function." That's the part of the genome that has a causal role and is conserved by selection at the organismal level.
Now look at the segment around 9 o'clock on their figure. That's DNA sequence that has a causal role and is also conserved. In this case the conservation is at a different level so it doesn't count as "function" by this definition. This is a case where a sequence seems to meet the selected-effect definition of function but it's being ruled out-of-bounds for other reasons. The best examples are active transposons and prophage such as integrated copies of bacteriophage lambda in E. coli.
This is a case where I prefer to stick to an unqualified selected-effect definition and call those sequences functional and not junk. (It's not clear whether the authors of those papers put them in the junk DNA category. I've asked them but they give equivocal answers.)
A sequence is conserved but not functional
Are there sequences that meet the selected-effect definition but don't have a biological function? Yes there are, but the border is fuzzy.
The problem here is operational rather than philosophical. Pseudogenes show clear evidence of sequence similarity when you compare different species but they are junk. You may argue this doesn't count because careful examination shows these sequences are drifting away from a common ancestor that once was an active gene. Eventually, their sequences will be indistinguishable from random sequence. True enough, but for now they are examples of sequences that meet the conservation criterion but they are junk.
There are other fuzzy examples. Many comparisons between genomes use small windows for their analysis. For example, they may look at 100 bp stretches along each of the genomes under comparison. Depending on their arbitrary cutoff, like 30% sequence similarity, they will detect many "conserved" sequences that are just due to chance. You need to be careful in assigning such sequences to the functional part of the genome. This isn't a problem with the definition, it's a problem with recognizing conservation.
De novo genes
De novo genes are new genes that have arisen in a particular lineage. There are problems identifying such genes because you have to determine whether they have a biological function before they count as genes. You can't use conservation as a criterion because, by definition, new genes aren't conserved. In this case you need to figure out whether the gene actually does something in spite of the fact it isn't conserved.
There's a nice review of the problems in the September issue of Nature Reviews: Genetics (McLysaght and Hurst, 2016). The number one problem is how to determine if a putative new gene is actually functional. This is a real problem since gene detection programs over-predict genes. Every new genomic sequence has dozens of potential new genes that have never been seen in any other species. These sequences are often referred to as ORFan genes because they have an open reading frame. The designation is unfortunate since they are actually potential or putative genes, not confirmed genes.
Most of these putative ORFan genes turn out to be false positives produced by the computer programs. That's why the number of putative ORFan genes drops precipitously as the draft genome sequence gets annotated.3 There are only a handful left in the human genome. Some of them are real and some of them are still ambiguous.
Since we're dealing with putative protein-coding genes, the best way to determine if they are real is to see if you can detect an mRNA and a protein. That eliminates most of the candidates. The next step is to find out if the (usually small) protein has a biological function or is just junk protein. That's much harder. These are all causal-role issues.
McLysaght and Hurst point out that it's still possible to look at conservation as a criterion by comparing sequences in a large number of individuals. If the sequence is under selection you expect to see less variation than you see in neighboring junk DNA. Unfortunately, these putative genes are all quite small so there's hardly any variation within the population.
There are no easy answers but trying to decide whether a putative de novo gene is functional highlights some of the problems with defining function.
UPDATE (Dec. 12, 2016)
I haven't made my personal position clear in this post. My views are the same as those I outlined in previous posts (see below). I agree with Sean Eddy (Eddy, 2013) when he says,
Attention focused on the squabbling more than the substance, and probably led some to wonder whether the arguments were just quibbling over the semantics of the word ‘function’.Here's what I said earlier ... my view hasn't changed.
Trying to conceptualize the forces that act on genome evolution is not just a matter of semantics.
Although I am going to quibble about the word “function” in this lengthy post, my main point is that the function wars are, for the most part, distracting and unproductive. We’re interested in the big picture—whether most of our genome is junk—and that’s not going to be resolved by settling on a definition of “function.” We have enough experience in biology to know that very few terms can be defined unambiguously (e.g. “gene,” “species”).I think of my view as being pragmatic (and scientific) as opposed to philosophical (and metaphysical). Look at the quibbling in the comments. Who the hell cares whether Ford Doolittle and Dan Graur have defined function in the same way as philosphers did several decades ago? The important point is whether pseudogenes are junk; whether most of intron sequences are junk; and whether fragments of transposons are junk.
I also said ...
My position if is that there's no simple definition of function but sequence conservation is a good proxy. It's theoretically possible to have selection for functional bulk DNA that doesn't depend on sequence but, so far, there are no believable hypothesis that make the case. It is wrong to arbitrarily DEFINE function in terms of selection (for sequence) because that rules out all bulk DNA hypotheses by fiat and that's not a good way to do science.But we can't work in a complete vacuum when it comes to function. There has to be some concept of what's functional and what's not. That's why I suggest the following as a "working definition."
So, we can adopt a working definition of function and junk based on whether or not deleting the DNA in question affects the survivability of the organism or its descendants. (Keeping in mind that there are minor exceptions).Keep in mind also, that we aren't really going to delete every bit of DNA to test whether it is junk DNA or not. A lot of the debate will be in the form of thought experiments where a likely conclusion will be suggested by what we already know about a specific sequence.
I do not intend this operational, working definition to be definitive or rigorous. Back off, philosophers. It's just a ball-park estimate of what it means to talk about junk DNA.
Function Wars
(My personal view of the meaning of function is described at the end of Part V.)
- On the Meaning of the Word "Function"
- The Function Wars: Part I
- The Function Wars: Part II
- The Function Wars: Part III
- The Function Wars: Part IV
- Restarting the function wars (The Function Wars Part V)
- The Function Wars Part VI: The problem with selected effect function
- The Function Wars Part VII: Function monism vs function pluralism
- The Function Wars Part VIII: Selected effect function and de novo genes
- The Function Wars Part IX: Stefan Linquist on Causal Role vs Selected Effect
- The Function Wars Part X: "Spam DNA"?
1. In this particular example, the conclusion is bolstered by the fact we have an explanation for the phenomenon; namely, randoms mutations in junk DNA that accidentally create a binding site. Such sites will soon be uncreated by additional mutations as the genome evolves.
2. Nobody disputes the evidence that most transposon-related sequence are defective or fragments of once-active transposons. Collectively they make up almost 50% of the human genome. They are clearly junk.
3. Unfortunately, most genomes never get past the draft stage so there are hundreds of genomes with large numbers of ORFan "genes" that will never be corrected. Intelligent Design Creationists love to focus on those genomes.
4. The underlined words ("in most eukaryotes") is an update. I added it after Georgi Marinov pointed out in the comments that there are some unusual species that have introns as small as 18-21 bp. Those species seem to have a different spliceosomal mechanism than the one found in most other eukaryotes. The important point is not the exact size of the minimal intron but the necessity of having a minimal size consisting of some sequence that's not conserved.
Doolittle, W.F., Brunet, T.D., Linquist, S., and Gregory, T.R. (2014) Distinguishing between “function” and “effect” in genome biology. Genome biology and evolution 6, 1234-1237. [doi: 10.1093/gbe/evu098]
Elliott, T. A., Linquist, S. and Gregory, T. R. (2014) Conceptual and empirical challenges of ascribing functions to transposable elements. The American naturalist 184:14-24. [doi: 10.1086/676588]
McLysaght, A., and Hurst, L.D. (2016) Open questions in the study of de novo genes: what, how and why. Nature Reviews Genetics, 17:567-578. [doi: 10.1038/nrg.2016.78]


92 comments :
I have to disagree with your examples of conserved sequences that are junk. Sequence similarity is not evidence of conservation. It's evidence of either conservation or short time since divergence. Conservation is (or should be, at least) defined as the result of evolution at lower than the neutral rate. A chimp sequence that's 98.7% similar to a human sequence is not conserved. A chimp sequence that's 99.5% similar (the average for protein-coding exons) is conserved. Nobody looks at human-chimp comparisons and claims a sequence is conserved because it's 98.7% similar to a human sequence.
I think the answer to your second question is "no", for any rational meaning of "conserved".
I don't understand your point? Did I ever say I didn't understand the criteria for sequence conservation?
When comparing genomes you will often run into stretches of sequence that are 70% identical. They might represent a functional gene in one genome and a pseudogene in another. We could quibble about the exact meaning of "conserved" in this case but it seems obvious to me that there has been selection in the past preventing those sequences from evolving at the neutral rate.
If you don't know that one of the sequences is a pseudogene then you will assume that both these regions are functional in the two genomes. That's a reasonable conclusion based on the selected-effect criterion.
You need to apply other tests to determine whether the sequences are junk.
This is a problem in whole genome analyses of sequence conservation. It demonstrates that the operational definition of conserved (high sequence similarity) doesn't necessarily correlate with function.
Another problem is when you look at recently duplicated genes in one species. You can be reasonably confident that one copy will eventually die by acquiring disabling mutations. It could be artificially knocked out to prove that it is redundant. It is conserved but effectively junk-in-waiting.
I was going to raise the same point, but since you already did, I'll just have to note that the rate we measure is the mean rate along lineages. This means that there is a chance to get false negatives, since a sequence that for some time has a rate above neutral (indicating darwinian selection) and then drops below neutral (indicating conservation) may well have a mean rate that is indistinguishable from neutral. It's another of the classical "failing to detect selection does not mean that there is no selection, just that we haven't detected it" cases. It's not going to make a big dent in our estimate of the amount of junk in our genomes (and there is a fair chance we could detect this if we used additional data - ancient DNA could be an option, as would comparisons between multiple human populations).
Larry, you didn't say you don't understand my point, but apparently you don't. Let me restate: % similarity != conservation. % similarity that's small compared to neutral expectation = conservation. 70% similarity between two species may or may not be considered conserved depending on the time since divergence, or more operationally by comparison to sequences we expect to have been evolving neutrally. 99% similarity may or not be considered conserved. There is no percentage similarity that counts as conserved other than by comparison to presumed neutrally evolving sequences.
And sure, there may be complications if there have been episodes of neutrality and selection at various points along a lineage. But that doesn't alter the point. "High sequence similarity" is not the criterion for conservation; it's high sequence similarity compared to neutral expectation. The second part is crucial.
...% similarity that's large compared to neutral expectation...
@John Harshman,
You are correct, but I still don't see how that invalidates the point that a sequence can be conserved but not be functional according to the causal-role definition.
I would think that a sequence that recently turned into a pseudogene can stil have high sequency similarity compared to the neutral expectation since it has experienced purifying selection for most of the time since divergence. I assumed this was the point that Larry tried getting across. Perhaps I misunderstand?
@John
Okay, now I get it. You assumed I was too stupid to know that recently diverged species show a high degree of sequence similarity across the entire genome just because there hasn't been enough time to diverge. That means the conclusion of "conservation" has to include a degree of sequence similarity that's greater than that expected by the null hypothesis.
I'm feeling a little insulted.
A small nitpick:
It seems to be about 50 bp, or enough to form the loop of RNA that's required to bring the 5′ and 3′ splice sites together in the spliceosome.
There are actually genomes with 20-30bp introns, and apparently even some with as low as 16bp
Regarding pseudogenes:
A recent pseudogene should have high (i.e. neutral) frequency of polymorphisms within the population, while functional protein coding genes should be constrained according to the same measure.
This is often forgotten in this discussion but actual population genomics is a real thing these days and it can be very useful with respect to such questions
References please. I can't imagine a spliceosomal intron of only 16 bp. I bet it's an artifact of overly imaginative alternative splicing predictions.
@Georgi,
Yes, you can tell if something is currently conserved or not by looking at polymorphisms within the population. However, as McLysaght and Hurst point out in their review, this only works with fairly large sequences in which case lack of conservation BETWEEN species should also be apparent.
The challenging cases for putative de novo genes are often quite small (ORF ~300 bp). There's not much polymorphism in such a short sequence. You need a massive number of genomes with highly accurate sequence in order to draw a conclusion from the absence of possible detrimental alleles.
It's a practical, not a theoretical, limitation.
Let's not lose site of the main point. The selected-effect criterion doesn't help you very much in deciding whether a new genes has a genuine biological function. You have to rely pretty much on causal-role effects.
I don't see why an intron couldn't be 16 bp if it isn't required that the RNA can fold back on itself. Wouldn't that depend on the mechanism by which it is spliced out?
I mean, a protein could at least in principle bind two sites close together on a piece of RNA and cut it out, like a restriction enzyme.
I can't give you a reference for the 16bp ones because I have seen a talk about that but no follow up paper (it's a weird ciliate species).
The smallest introns other than that are in nucleomorphs:
https://www.ncbi.nlm.nih.gov/pubmed/19380463
They are admittedly an outlier because they're not autonomous nuclei, but they are still eukarotic nuclei.
But free-living eukaryotes can have very small introns too. Again, ciliates are the prime example:
https://www.ncbi.nlm.nih.gov/pubmed/19061489
https://www.ncbi.nlm.nih.gov/pubmed/8165136
https://www.ncbi.nlm.nih.gov/pubmed/17086204
Also, the ultratiny introns are not predictions, but derived from RNA-seq data
We will have many millions of human genomes in the coming decades though. And after a hundred years or so of sequencing everyone at birth, it should become apparent what is constrained and what is not as every nucleotide will have had ample opportunity to be mutated multiple times.
I enjoy most of the nitpicking but it's also very frustrating. I spend most of my days trying to write a book that will be comprehensible to a large audience of non-experts. It's not easy.
Others have published books on junk DNA but I suspect they didn't have the same problems I'm having. They didn't struggle with how to describe complicated arguments and heated controversy among scientists because they were mostly writing from ignorance of those issues.
It's much easier to write a book if you think you know all the answers and you can ignore everyone who disagrees with your viewpoint. That's how Nessa Carey manages to publish a new book every year.
I'm constantly struggling with something that Richard Dawkins calls "yes-buttery." It's the response you often resort to when you read something that's not quite right. You say to yourself; "Yes, but there are some introns smaller than 50 bp."
Too much yes-buttery and you lose the respect of your knowledgeable readers. But there's a trade-off. In order to cover all the exceptions and quibbles you lose your other readers.
I'm trying to partially solve the problem by adding footnotes. For example, I say a gene is a "DNA sequence that's transcribed" then I add a footnote for John Harshman and Simon Gunkel to make sure they know that I'm aware of RNA genomes! :-)
The other trick is to insert qualifiers like, "As a general rule, introns have to be at least 50 bp in order to be efficiently spliced." However, too much of this kind of writing makes the book difficult to read.
Sometimes there's just no way to avoid complications. For example, I've decided to insist on my definition of a gene and on the fact there are noncoding genes as well as genes that encode proteins. I'm trying not to equate "genes" with "protein-coding genes" and that means I often have to use the correct phrase (protein-coding gene) where others are getting away with just "gene." You can bet that if I slip up there will be many Sandwalk readers who will say, "yes, but ..."
These issues and problems were very much on my mind last week when I wrote about alternative splicing. It would have been so easy to mouth the standard platitudes about alternative splicing and how it creates 100,000 proteins from only 20,000 genes. That's what Nessa Carey and John Parrington did in their books on junk DNA.
It would also have been easy to simply declare that the vast majority of human genes make a single protein and just leave it at that. That's not really an option since the popular literature and the scientific literature are full of stories about the importance of alternative splicing. In order to present my view I have to also show why those stories are wrong.
So, I have five complicated pages on the problem with difficult hard-to-follow arguments about the quality of the scientific literature. Other authors get way with just one or two sentences extolling the virtues of alternatively spliced human genes and how they explain complexity.
I even quote Nessa Carey to illustrate the problem. In her book Junk DNA she says on page 238, "At least 70% of human genes have been shown to create at least two proteins."
She treats alternative splicing as just a simple fact she needs to tell her readers about.
Georgi says,
"Also, the ultratiny introns are not predictions, but derived from RNA-seq data."
Well that settles it then. We all know how reliable RNA-seq data is at predicting genuine functional transcripts. :-)
It wasn't my intention to sabotage things with nit-picking, I just noticed that the sentence wasn't correct and my instinct to always bring in the additional information when it would improve the factual accuracy of the text kicked in.
I fully understand your challenge. BTW, it is not only easier to write books if you do not know too much, it is also a lot easier to do research. With all the consequences, and they are not always negative (sometimes that is empowering and enables people to do things that otherwise they would not dare follow to completion)
Well that settles it then. We all know how reliable RNA-seq data is at predicting genuine functional transcripts. :-)
It's not about prediction, it's just that when you map reads (or assemble transcripts and map those) to the genome, this is what you see. And that the sequence of proteins only makes sense with introns that are not a multiple of 3.
Having a million high quality genomes could work as long as long as you can identify recessive detrimental alleles.
I bet Michael Lynch has thought about this problem. Given a coding sequence of 300 bp that only exists in humans, how many genomes would you have to sequence in order to detect a significant number of polymorphisms? How many potentially detrimental alleles would you need in order to be confident that the coding region wasn't functional?
Does anyone know how many loss-of-function alleles are present in the human population for an average protein-coding gene? That's the background level you need to subtract in order to determine whether a potential gene has a function.
Even then, it gets complicated. In my book I discuss the gene for N-acetylgalactosaminyltransferase. There are millions of people who are homozygous for loss-of-function alleles of this gene. They have O-type blood because this is the gene that determines ABO blood groups.
If you have O-type blood group then that gene in your genome is a pseudogene. It is junk DNA. Does that mean the gene is also junk in people with blood types A or B even though it makes a functional enzyme?
How does the selected-effect criterion work in this case?
Larry,
Don't feel insulted. I was only criticizing what you wrote, which is the only way I can infer what you think. The fact is that we've argued about this before. Why not change what you say so that your meaning is clear, rather than attacking me for taking you at your word?
Thanks Larry for your interesting exposition of difficulties in determining whether or not a certain DNA segment is functional!
You conclude by saying that "trying to decide whether a putative de novo gene is functional highlights some of the problems with defining function."
I have the impression that both you and McLysaght & Hurst implicitly employ the notion of function as biological role, that is, roughly speaking, a definition of function as the role of a part or activity of an organism in maintaining that organism's capacity to grow, develop, survive and reproduce. If so, the difficulties you highlight aren't difficulties with the *definition* of function but difficulties with the operationalization of the notion of function as biological role in the context of research into the functionality of DNA segments. In other words: the problems with which you are concerned is the problem how to find workable criteria that indicate whether or not a putative de novo gene has a function as biological role.
If so, there is no need to revive boring wars over the meaning of the word "function". Instead, you can focus on the development of suitable criteria for recognizing functionality in different contexts (which is in effect what McLysaght & Hurst do). A much more interesting activity!
Does that make sense?
No, it doesn't make sense. You are correct to assume I'm interested in detecting function and, by implication, junk DNA. However, it seems to me that defining the word "function" is incredibly important for this task. After all, it's the problem of definitions that got us into the ENCODE mess a few years ago.
I think conservation is our best bet for detecting function in the vast majority of cases (selected-effect). I'm happy to support that definition over all others, such as causal-role.
I just want to make sure we can all agree on how to apply that definition (i.e. active transposons?). I also want to make sure we don't get carried away with the best definition and start thinking it's the only way to confirm function. That's why I brought up some tough examples like de novo genes and bulk DNA.
The function wars are, indeed, boring to most people. That's partly because they usually don't deal with interesting and challenging examples and partly because they often degenerate into philosophical mumbo-jumbo.
I agree with Gould that something other than semantics is usually at stake. That doesn't mean that semantics isn't important was well.
John, you say, "Why not change what you say so that your meaning is clear, rather than attacking me for taking you at your word?"
Okay, let's give it a try. Here's what I wrote.
There are other fuzzy examples. Many comparisons between genomes use small windows for their analysis. For example, they may look at 100 bp stretches along each of the genomes under comparison. Depending on their arbitrary cutoff, like 30% sequence similarity, they will detect many "conserved" sequences that are just due to chance. You need to be careful in assigning such sequences to the functional part of the genome. This isn't a problem with the definition, it's a problem with recognizing conservation.
I assume you object to the 30% example because it only applies to genomes that are so distantly related that the overall sequence similarity should be 25% just by chance. I admit I was thinking of protein sequences when I wrote that so I should have used a different example. I really don't think you had trouble understanding my meaning. And I really don't think you believe that I'm too stupid to understand sequence similarity.
What you might have said was, "Larry, I know you understand this stuff but some people may misinterpret what you wrote. Why not say ...?"
Instead you choose another approach. Whatever. How about I edit my words to say ...
For example, they may look at 100 bp stretches along each of the genomes under comparison. Depending on their arbitrary cutoff, like 70% sequence similarity in distantly related species, they will detect many blocks of "conserved" sequences that are just due to chance.
Larry: "it seems to me that defining the word "function" is incredibly important for this task."
I agree. My point is that there is a good definition of function that serves to glue biology together, namely function as biological role. In case of DNA segments the best operationalization of that notion is sequence conservation. That criterion does not apply when talking about the function of hearts, but there are other ways to operationalize the notion of function as biological role in that case. However, if function is *defined* in terms of sequence conservation this would mean functional morphologists who say that the heart is the source of energy of the circulatory system and molecular biologists who say that the function of a certain gene is specify the amino-acid sequence of a certain protein are talking about different kinds of function and that seems absurd. Also note, that sequence conservation is an indication that a certain segment has a function, but a definition of function in terms of sequence function is of no help in determining what that function is. For that reason, I belief there is no serious problem in defining function, but that there are serious difficulties in the operationalization of that notion in the case of DNA segments.
As another example of "conserved but not functional", I guess you could invoke telomere repeat units. The sequence is conserved because of the mechanism that synthesises them, but any individual repeat unit can't really be said to have a function: they only have a function in aggregate. It's the flip side of the "bulk DNA" theory you discuss.
In regard to the wider question of function, I think that the definition of "function" is context sensitive, and that in some contexts it is overly restrictive to use the selected-effect definition.
There are genes I bear - or more accurately genetic variants - that make me phenotypically different from my brother. Hairy toes, monobrow versus unibrow, blobby nose versus pert, what have you. The traits are unmistakably heritable, and you can trace them over generations in the family album. And yet, it is vanishingly unlikely that there is natural selection on monobrows, or blobby noses. These are selectively neutral phenotypes mediated by selectively neutral genetic variation.
One can generalise this. Any DNA variation that is present at high frequency in the human population - i.e. pretty much everything that makes us all genetically and phenotypically different from each other (twins aside) - is likely to be selectively neutral. If it were strongly selected, then the variation would already have been removed! It seems perverse to dismiss the DNA elements responsible for all individual human phenotypic variation as "functionless", and yet that's what the selected-role definition implies.
This variation can be highly significant. Genetic diseases with onset after reproductive age are often not efficiently selected against. The causative variants will be selectively neutral or nearly so. I think that in the context of genetic medicine it is perfectly legitimate to say that a given stretch of DNA is 'functional' because it binds transcription factor X, which causes the expression of gene Y, which ultimately prevents disease Z. If you delete that bit of DNA, it can no longer bind X, so Y is no longer expressed and you suffer from disease Z. This holds true irrespective of whether disease Z affects reproductive success and thereby leads to sequence conservation of the underlying causative DNA sequence.
Ultimately it is a what-versus-why distinction. The causal-role definition tells you what a gene does, the selective-effect definition tells you why it does it. Both can be called "function" in different contexts.
Addendum: the strength of selection on a given variant depends on population size. Does it really make sense to say that a given stretch of DNA is "functional" if the population size is greater than a given threshold, but loses its function if the population size drops low enough that selection is no longer effective at that locus?
If a given transcription factor binding site in my cells is functional today, can it be functionless tomorrow if J Random Dictator goes mad and triggers a nuclear war?
I think it would be better if you just explain up front that "conserved" should mean "conserved relative to neutral expectation", then explain that there's expected variance to be accounted for too. And then in the examples, including hypothetical ones, you should mention the expectation.
I don't agree that any of your examples counts as "conserved but not functional". The closest you get is "conserved until fairly recently but no longer functional", which I would count as different.
Larry, I know you understand this stuff, but some people may misinterpret what you wrote. I think we still have a disagreement about whether "conserved" ha a strictly operational meaning or whether it refers to underlying processes, i.e. whether it just means a certain amount of relative sequence similarity or whether the similarity is merely used as an attempt to diagnose sequences undergoing purifying selection. Or perhaps it has either of these meanings depending on context.
Larry: "After all, it's the problem of definitions that got us into the ENCODE mess a few years ago."
I don't think so.
ENCODE explicitly defined its notion of function:
"Operationally, we define a functional element as a discrete genome segment that encodes a defined product (for example, protein or non-coding RNA) or displays a reproducible biochemical signature (for example, protein binding, or a specific chromatin structure)"
As Germain et al. point out in their excellent "Junk or functional DNA?" (Biology & Philosophy 29:807–831, 2014) ENCODE aimed to "identifiy relevant activities—that is, activities likely to make a relevant difference to some phenomena scientists are likely to care about." It seems to me that in view of that goal, ENCODE's definition of function as biochemical activity is appropriate.
So the mess didn't occur because ENCODE used the wrong notion of function, but because they failed to point out that their notion of function as activity is different from the notion of function as biological role used to define junk DNA.
And the mess was increased by papers such as Graur et al.'s "On the Immortality of Television Sets" and Doolittle et al.'s "Is junk DNA bunk?" which, instead of pointing out that 'having a function as activity' does not imply 'having a function as biological role', blamed ENCODE for not using the teleological notion of function as selected effect developed by Millikan and Neander to solve some vexing problems in the philosophy of mind and language. Both papers confuse the notion of function as selected effect with the notion of function as biological role, thus creating a needles function war, where semantic clarification such as that of Germain et al. would have been a more appropriate and fruitful response.
I'm just curios Larry; do you believe your hair has a function?
I know it is kind of odd question but I just would like to get your prospective on simple thing like that. I hope you don't mind?
I don't really know what you mean by this question about courts and lawyers. Are you asking whether I agree with Larry? Whether I think it's my duty to defend Larry? I think Larry is mostly right. We disagree on only a couple of points. I agree that conservation is the best, or at least most available, evidence for function. We may differ slightly on what "conservation" means. I also would not agree that transposable elements are functional; their sequences certainly aren't conserved by selection. But that's all I can think of at the moment.
Are you one of the resident creationists under a new name?
John Harshman,
You kick my brain into a higher gear most of the times you comment! Thanks.
I have question for you. Hope you don't mind.
If a judicial court existed for scientists and you were the lawyer defending Larry's view of junk-DNA, would you take the job?
I'm not going to do links and shit. I just want your truthful unbiased opinion without sacrificing your future if possible I hope
Thanks John.
I'm not a creationist.
You explained yourself enough.
Thanks John.
I'm not a creationist.
You explained yourself enough.
Yeah, we have to look for a particular function in humans, because the fact all mammals have hair wouldn't indicate anything about heritable genetic characteristics, would it?
Of course you aren't. You just play one on the internet.
What he said. I am a big fan of Arno's publications on this - if more people had read them, instead of sticking to the dichotomy between selected effect and casual role we might have avoided the semantic debate in favour of a substantive scientific one.
I'd take that job.
"So the mess didn't occur because ENCODE used the wrong notion of function, but because they failed to point out that their notion of function as activity is different from the notion of function as biological role used to define junk DNA."
It seems to me these two errors are functionally equivalent in that they lead to the same mess: Lots of upcoming biologists and the general public now thinks it has been proven that the entire genome is selected-effect functional. Even some molecular biologists who should know better think they did (e.g. John Mattick).
Arno, do you think ENCODE's failure to point out the difference between function as activity and function as biological role was merely an oversight, or that they wanted to have articles/headlines saying the vast majority of the genome is "functional"?
I do take some comfort in the fact that so many creationists are embarrassed to admit that is what they are. They are not quite as shameless as they usually appear.
Larry,
I am a regular reader of your very interesting blog. As an evolutionary biologist now mostly involved in lecturing, I look forward for your book. As an author of a small volume about evolution in german I would say that it is much better to tell a story than to explain difficulties. Some citations of papers are important to Illuminate mechanisms or to evaluate hypotheses, however, generally, dry facts are boring. If I had problems to support or disprove hypotheses, I skipped this topic altogether. Every eukaryote seems to have a specific intron size which represent the modus and also (nearly) the minimum. In Drosophila melanogaster, it is 60 bp. For the lay reader, however, it is only important to explain the well-supported hypothesis of Martin and Koonin about the initial evolution of introns
http://www.nature.com/nature/journal/v440/n7080/full/nature04531.html
which points to a tight interaction of endosymbiosis, parasitic elements of the genome and drift. This story would be enlightened, not the lower limit of intron size.
I unterstand function in the sense of evolutionary conservation because this means that the corresponding sequence takes part in the reproduction of an evolutionary unit. This has not to be an organism, as viruses and transposons are also conserved. Retrotransposons would not be conserved on a specific genomic site but by transposition. Such elements are functional for themself, not in relation to the host. Telomers are functional for the organism, irresponsible of the mechanism of their sequence conservation, which is only indirectly related to selection, to some extent similar to the concerted evolution of ribosomal RNA loci. In contrast, if, indeed, some genome sequences effects only the health of such organisms positively which are no longer able to reproduce: Such sequences should not be called functional.
Is this conclusive?
Arno:
"It seems to me that in view of that goal, ENCODE's definition of function as biochemical activity is appropriate"
I think the ENCODE definition of function is obviously muddled. It was wasn't they wouldn't have made a very transparent mistake. They initially said that by their criterion 80% of the genome is functional. But any bright high school student could have pointed out that by their criterion 100% of the genome is functional since every single nucleotide is read and copied by DNA polymerase
Arno,
"It seems to me that in view of that goal, ENCODE's definition of function as biochemical activity is appropriate."
The obvious next question is to ask what that definition is appropriate for. Is it appropriate for describing functions that are impact fitness? I think it is quite obvious that the ENCODE is not appropriate in this context. A large portion of the DNA sequences labeled as functional by ENCODE using that definition could be removed from the human genome and it wouldn't affect fitness one iota.
To use an analogy, imagine your TV stopped working just a day after you bought it. You take it back to the store, but they refuse to refund your money because they claim your TV is still functional. You ask the store manager how your TV is still functional, and he says that it is still able to grab dust from the environment and adhere it to the surface of the TV. That would fit ENCODE's definition of functional, because the TV is still interacting with the environment and changing the chemistry of its surroundings.
ENCODE made the mistake of confusing "does something" with "functional". Those are not the same thing.
Peter,
"And yet, it is vanishingly unlikely that there is natural selection on monobrows, or blobby noses. These are selectively neutral phenotypes mediated by selectively neutral genetic variation."
There are also variants of those same genes that are detrimental. While a monobrow may be selectively neutral, mutations in those same genes that result in a malformed cranium which doesn't allow for proper brain development will be selected against. It is the selection against these detrimental mutations that results in the observation of sequence conservation.
The onion shirt is quite snarky. I like it. You could also have another shirt with the words, "Keep calm and ask about the bladderwort". This plucky little plant has about the same number of genes as any other plant or eukaryote, but its genome is only 82 million bases.
The plants at both ends of the genome size spectrum pose interesting questions for those trying to argue for function in large portions of a genome.
Arno Wouters said,
So the mess didn't occur because ENCODE used the wrong notion of function, but because they failed to point out that their notion of function as activity is different from the notion of function as biological role used to define junk DNA.
That's a very serious misreading of what happened. The major ENCODE Consortium leaders are on record as saying that their data disproves junk DNA. Their definition of "function" was fully intended to be used as evidence against junk DNA. Many of those leaders continue to be skeptical of junk DNA because they think most of the genome is functional (i.e. biological function).
They actually believe that pervasive transcription means biological function. They actually believe that all those millions of transcription factors binding sites and DNAse I hypersensitive sites are involved in gene regulation. Read the papers.
What did the ENCODE Consortium say in 2012?
How does Nature deal with the ENCODE publicity hype that it created?
Science still doesn't get it
The popular press was full of articles announcing the death of junk DNA but, more importantly, so were the science journals, led by Nature and Science. You did not see any outrage from the ENCODE Consortium leaders about this so-called "misrepresentation" of their data. That's because those leaders really though their definition of function ruled out junk DNA.
It wasn't until eighteen months later that they started to complain about the media "misrepresenting" their conclusions (e.g. Kellis et al., 2013). If you really think ENCODE leaders didn't mean to disprove junk DNA then I challenge you to find a single example of a major ENCODE Consortium group leader who spoke out at the time (2012) to complain about the coverage and the widespread publicity about the death of junk DNA.
Check out the links above and watch the video where Ewan Birney appears with a Nature editor who says, "The striking overall result that the ENCODE project reports is that they can assign a function, a biochemical function, to 80% of the human genome. The reason why this is striking is because, not such a long time ago, we still considered that the vast proportion of the human genome was simply junk because we know that it's only 3% that encodes proteins."
Does that sound like someone who is making a distinction between the ENCODE definition of function and "not-junk"?
One of the "News & Views" articles that accompanied the ENCODE papers in Nature (Ecker et al. (2012) says,
One of the more remarkable findings described in the consortium's 'entrée' paper (page 57)2 is that 80% of the genome contains elements linked to biochemical functions, dispatching the widely held view that the human genome is mostly 'junk DNA'. The authors report that the space between genes is filled with enhancers (regulatory DNA elements), promoters (the sites at which DNA's transcription into RNA is initiated) and numerous previously overlooked regions that encode RNA transcripts that are not translated into proteins but might have regulatory roles.
Did you see any ENCODE leaders complaining about this "misrepresentation"? No you didn't. That's because they thought their definition of function defined a biological function that ruled out junk DNA.
They were wrong. They don't get to escape being wrong by promoting some revisionist history.
The problem is that there isn't just one hair gene. Hair is the result of an interaction between many, many genes which each have their own role within the development of many other morphological features. Hair could be a spandrel hitching a ride with some other selected function (e.g. skin development).
Arno Wouters says,
As Germain et al. point out in their excellent "Junk or functional DNA?" (Biology & Philosophy 29:807–831, 2014) ENCODE aimed to "identifiy relevant activities—that is, activities likely to make a relevant difference to some phenomena scientists are likely to care about."
I surprised that you would promote such a bad paper. The goal of Germain et al. (2014) is to promote their own views of function and junk and they do this, in part, by retroactively reinterpreting what the ENCODE leaders "really" meant two years earlier.
In fairness, Germain et al. do hint that ENCODE may not have behaved properly,
In the headlines of The Washington Post one could read "Junk DNA concept debunked by the new analysis of human genome." The ENCODE Consortium, which made little attempt to clarify their point, was accused of contributing to this misunderstanding.
But they certainly did NOT contribute to the "misunderstanding." They agreed with the headlines. They didn't think it was a misunderstanding at all.
The paper by Germain et al. rambles on about various definitions of function and various definitions of junk. They focus on some mystical relevance related to something called "biomedical research." They say in the abstract,
... we argue that ENCODE's controversial claim of functionality should be interpreted as saying that 80% of the genome is engaging in relevant biochemical activities and is very likely to have a causal role in phenomena deemed relevant to biomedical research.
What the heck does that mean? About 50% of our genome consists of fragments of defective transposons. In what way are those sequences "relevant to biomedical research" other than proving they are junk DNA?
More than 20% of our genome consists of intron sequences that are not conserved and do not serve any biological role other than taking up space between splice sites. In what way are these sequences "relevant to biomedical research"?
I claim that junk DNA is non-functional DNA. That means it makes no significant contribution to biological function. It could be deleted without affecting the organism or its descendants.
Here's what Germain et al. say about such a definition.
Finally, if junk DNA is instead taken to mean DNA that does not make a significant difference to biological phenomena of interest to biomedical research, then ENCODE's claim of functionality of the genome does conflict with the view that most of our genome is junk.
So, ENCODE's claim of function DOES conflict with my view of junk DNA according to Germain et al.!
Paul Griffiths says,
What he said. I am a big fan of Arno's publications on this - if more people had read them, instead of sticking to the dichotomy between selected effect and casual role we might have avoided the semantic debate in favour of a substantive scientific one.
I haven't read his papers on function because they don't deal with the practical issues of defining junk DNA and function in the genome.
Perhaps you could enlighten us with a short description of "function" that helps us answer questions about the amount of junk DNA in our genome?
I recall a brief discussion we had with Karola Stotz in London. She claimed that most human genes makes several different proteins as a result of alternative splicing. Do philosophers have a solid definition of "function" that helps us determine whether she is correct or not? How do philosophers know that all those transcript variants have a function? What criteria should we use to answer the question?
Your talk was about information in the genome. I assume that's related to function in some way. But you only talked about genes, if I recall correctly.
Do origins of replication and centromeres contain information? Do they have a function? Have you got a substantive scientific definition of function that covers these sequences?
I guess you are both clueless about the "possible function" of hair as a whole.
How about eyebrows specifically? What possible function could they have from evolutionary prospective obviously?
Don't bite your nails just yet!
John Harshmn,
Who do YOU think a creationist is? I don't want the popular definition. I'd like YOU to tell me how you view or interpret that someone, like myself, is a creationist.
In the 1980s there were "scientific creationists" who were happy to claim the word. Since then the public has had a more and more negative impression of creationists, so now actual creationists are insisting that they are not creationists but "intelligent design theorists". Some of those intelligent design theorists just happen, by pure coincidence, to think that the universe is 6,000 years old.
Don,
You should read Larry's post on the problems with dogmatic adaptionist worldviews. Also, read up on biological spandrels:
https://en.wikipedia.org/wiki/Spandrel_(biology)
Mikkel, judmarc, and lantog, I don't want to become involved in function wars, I am not interested in a game of shame and blame, and I am not going to speculate about the causes of ENCODE's failure to explicitly distinguish between function as activity and function as biological role.
My interest is in clearing up the mess created by ENCODE's failure followed by the subsequent introduction of the philosophical and teleological notion of function as selected effect by some of its critics (who confuse it with function as biological role). I believe that the first step needed to clear up this mess is to clearly distinguish function as activity, function as biological role and function as selected effect (there is a fourth notion of function, namely function as biological advantage but as far as I can see, this notion is not (yet?) relevant to the confusion).
I belief this work has been done already: I distinguish these notions in my Four Notions of Biological Function (2003) and Germain, Ratti and Boem apply them to clear up the ENCODE mess in their Junk or functional DNA? (2014).
I believe that Graur's (and others)' identification of ENCODE's notion of function as biochemical activity with Cummins' (1975) notion of function (often, but in my view erroneously, called 'causal role function') is very confused. ENCODE was concerned with the question 'what does it do?', whereas Cummins' functions are answers to the question 'how is it useful (in the sense of how does it contribute to a capacity or ability of interest)?' Hurst's (2013) claim that following a collision between a car and a pedestrian, a car’s bonnet could be ascribed the Cummins function of projecting a pedestrian many meters, is plainly absurd (see Carl Craver's "Role Functions, Mechanisms, and Hierarchy" - Philosophy of Science 68: 53-74, 2001). There can be no such things as Cummins functions *per se* for Cummins functions are by definition relative to an analytic account and a capacity of interest.
Graur's (and others) identification of biological function with Millikan's (1989) or Neander's (1991) notion of function (it is not clear with which one he identifies it) is confused too. Here is Millikan's (1989) definition:
"Putting things very roughly, for an item A to have a function F as a "proper function", it is necessary (and close to sufficient) that one of these two conditions should hold. (1) A originated as a "reproduction" (to give one example, as a copy, or a copy of a copy) of some prior item or items that, due in part to possession of the properties reproduced, have actually performed F in the past, and A exists because (causally historically because) of this or these performances. (2) … "
And this is Graur's:
"Accordingly, for a trait, T, to have a selected-effect function, F, it is necessary and (almost) sufficient that the following two conditions hold: (1) T originated as a “reproduction” (a copy or a copy of a copy) of some prior trait that performed F (or some function similar to F, say F′) in the past, and (2) T exists because of F (Millikan 1989). In other words, the selected-effect function of a trait is the effect for which the trait was selected and/or by which it is maintained."
Millikan's definition is historical through and through: an item's function (e.g. the function of Bello's heart) is completely determined by the achievements of that item's ancestors (e.g. the achievements of Bello's ancestors' hearts) and, hence, independent of that item's current features. If Bello has a heart because the hearts of Bello's ancestors pumped blood, it is the Millikan function of Bello's heart to pump blood, whether or not Bello's heart actually pumps blood, whether or not Bello's heart is capable of pumping blood, whether or not pumping blood is favorable to Bello, Bello's reproductive succes or the continued existence of Bello's heart, and whether or not pumping blood helps to maintain the trait.
Graur's definition on the other hand allows traits to have functions not only on the basis of their ancestral performance, but also on the basis of their current or even future performance: the selected-effect function of a trait is the effect for which the trait was selected and/or by which it is maintained. This is Bigelow & Pargetter's 'forward looking' notion of function, criticized by Millikan in another paper that appeared in 1989: 'An Ambiguity in the Notion of "Function"' (Biology and Philosophy, 4: 172-176) for being 'notoriously indeterminate'.
There are many more problems with Graur's distortion of Millikan's and Neander's notion of function as selected effect but I hope this suffices to convince you that Graur's dichotomy creates more heat than light.
Hm. I thought I posted my response in two parts after lantog's and long before Eric's but I had problems with my internet connections and apparently my response was posted just now. I need some sleep now, but I will try to find time to respond to the other comments directed at me sometime tomorrow.
Don, why don't you save all some trouble by telling us what you are? And who are you really? Are you saying you've never commented on Sandwalk before appearing as "Don Quixote"?
Don Quixote bristles at being called a creationist, yet expects Larry to provide adaptationist answers to his questions. No one can combine ignorance and hypocrisy quite like a creationist can
Larry says (about my reference to Germain et al.'s "Junk of functional DNA"?
I [am] surprised that you would promote such a bad paper. The goal of Germain et al. (2014) is to promote their own views of function and junk and they do this, in part, by retroactively reinterpreting what the ENCODE leaders "really" meant two years earlier.
I don't see why it would be a problem that Germain et al. promote their own views of function.
I fail to see that they are, 'retroactively reinterpreting what the ENCODE leaders "really" meant two years earlier'. In my view they interpret a claim in ENCODE's introductory paper, rather then the motives of the people who put that claim there and the appropriatenes of that behavior of ENCODE's leaders. Why would that make the paper bad?
They note that one of the reasons why the debate got so heated lies in the perception that the ENCODE consortium "was perceived as deliberately encouraging misinterpretations to artificially boost its success (and hence future funding)". They do not argue for or against that interpretation of ENCODE's motives, but, as you said, they give the impression that ENCODE may not have behaved properly. What's wrong with that?
They do interpret the motives of ENCODE's **critics** were they they identify "the worry that ENCODE’s notion of function will lead to a multiplication of irrelevant functions" as those critic's main worry and suggest that the critics invoke the notion of selected-effect function as bulwark against those irrelevant functions. Do you you think they err in this interpretation of the criticism that ENCODE used the wrong notion of function?
Next they announce their argument: "We argue that the selection- criterion is only one way (and an imperfect one) to identify relevant functions, and that ENCODE’s notion of function is suited to the aims and scope of the project."
I really can't see what's wrong with this setup.
Larry about the ENCODE leaders:
The major ENCODE Consortium leaders are on record as saying that their data disproves junk DNA. … Many of those leaders continue to be skeptical of junk DNA because they think most of the genome is functional (i.e. biological function). They actually believe that pervasive transcription means biological function. They actually believe that all those millions of transcription factors binding sites and DNAse I hypersensitive sites are involved in gene regulation.
I know, I know!
My point is that it is much more appropriate and much less confusing to respond by starting a debate about the question whether or not pervasive transcription indicates involvement in gene regulation than by starting a war about the notion of function.
Their definition of "function" was fully intended to be used as evidence against junk DNA
It is not clear to me how a definition can be used as evidence, but I would think that it is much more appropriate and much less confusing to point out that given ENCODE's definition of function, it is not sufficient to show that something has an ENCODE function in order to conclude that it has a biological role, than to blame ENCODE for using the wrong definition of function.
They were wrong
Perhaps, but to show that they were wrong we need to discuss the evidence and the conclusions that can be drawn from that evidence, rather than confused complaints about the notion of function they used. That is, we need a substantive scientific debate (hi Paul! Thanks for you support!) instead of a quasi-philosophical one.
I guess I have been clear enough by now.
The function of eyebrows and eyelashes is to house Demodex, tiny mites that live in the follicles. Of course.
One more attempt to make myself clear.
Larry said "I claim that junk DNA is non-functional DNA. That means it makes no significant contribution to biological function. It could be deleted without affecting the organism or its descendants."
So you use the term 'biological function' in a sense that implies "it could be deleted without affecting the organism or its descendants". In my view this clearly indicates that you use the phrase 'biological function' in the sense of, what I call, function as biological role. But let us ignore that for the moment and agree on the following language:
* let's call your notion of biological function 'Moran function'
* let's define 'Moran junk' as 'doesn't have a Moran function'
In these terms you claim that most DNA is Moran junk, in other words that most DNA has no Moran function. Right?
ENCODE defines 'a functional segment' as 'a segment that "encodes a defined product (for example, protein or non-coding RNA) or displays a reproducible biochemical signature (for example, protein binding, or a specific chromatin structure)"'. Let's call that an ENCODE function.
ENCODE claims that they have been able to assign an ENCODE function to 80% of the human genome. Right?
ENCODE does *not* define 'junk DNA' but you and many others have the impression that its leaders are trying to push the idea that ENCODE's results show that most DNA has a Moran function and is hence Moran junk. Right?
You disagree with that idea. Right?
Now, the claim that a certain stretch of DNA (let's call that stretch 'Bello') has an ENCODE function is logically compatible with the claim that that stretch of DNA (Bello) doesn't have a Moran function. Right?
How to proceed in this situation?
One way (the one I advocate) is to raise the following substantial issue: does the finding that 80% of our DNA has an ENCODE function imply that those 80% have a Moran function and hence aren't Moran junk?
This would mean of course that you clearly define the notion of Moran function (I suggest that you define it along the lines of my function as biological role – but that is not the issue I am pressing now!).
The other way (taken by Graur et al. and Doolittle et al, and, apparently, advocated by you) is to start a war on the notion of function by blaming ENCODE for using the notion of ENCODE function rather than that of Moran function (or instead of Graur function or Doolittle function).
The latter response is, in my view, not constructive.
One reason is that this response hides substantive scientific issues (such as disagreements about research goals, methodologies, the conclusions that can be drawn from ENCODEs findings and the way to proceed in the future) under the cloak of a philosophical quibble about the right notion of function.
Another is that Graur et al.'s and Doolittle's et al.'s notions of function are very confused. They force a dichotomous framework that has been fruitful in philosophy of mind and language upon a discussion in biology in which quite different uses of 'function' surface. To do so they secretly modify both Cummins' and Millikan and Neander's notions of function in a way that makes their notions incoherent and, hence, useless.
With all due respect, I do not think you are making a very convincing case. ENCODE's own written statements at the time do not distinguish between "Moran function" and "ENCODE function" and, in fact, are worded so a to imply that the latter demonstrates the former.
http://sandwalk.blogspot.ca/2014/05/what-did-encode-consortium-say-in-2012.html
I fail to see why you would take Larry, Dan Graur and others to task for doing exactly what you say they should have: Pointing out that the two definitions of "function" are not synonymous. It was ENCODE who failed to make that clear.
Arno, it is laudable to attempt to put a debate on a substantive scientific footing. However, since the vast majority of the scientists concerned with the debate, as well as those members of the public interested in the debate, use the word, "function," I think a project that relies on everyone to stop using this word, however confused/confusing and controversial, is probably a non-starter.
I therefore tend to agree with Dr. Moran that as long as people are going to use the word, and as long as having a "function" means "useful for some purpose" in ordinary parlance, then it's a good idea to actually concentrate on the useful purposes to which a bit of DNA has been or is being put.
I also think that part of what you deem "confusion" on the part of Graur and Millikan is nothing more than a recognition that, along with other processes at play in evolution, usefulness is evaluated on an ongoing basis via selection. Thus we may have sequences of DNA in our genome that don't serve (as sequence) a particular purpose (most of it); we may have sequences that once served a useful purpose but don't any longer; sequences that continue to serve a useful purpose; or de novo sequences that are serving new (in evolutionary time) useful purposes. I don't see anything particularly confusing about this.
Of course this is terribly oversimplified, but after all, it has to fit in the space of a comment on a blog! :-)
No one can combine ignorance and hypocrisy quite like a creationist can
Indeed, like the fact that if hair has a particular function, humans, elephants, and some other animals apparently need it less than animals such as dogs, cats, other apes besides humans, etc. Or it could be evolutionary accident - founder effects or other sorts of genetic drift....
Why did some dinosaurs suddenly "need" feathers? 'Cause they were gonna turn into eagles and peacocks in a few tens of millions of years? And why did platypuses decide hair was better than duck's feathers? Didn't they wanna fly too?
Go ahead, DQ, keep thinking of evolution in terms of "just so" stories, it will get you far, I'm sure.
Here's one for you - what's the purpose of nipples on guys?
Larry to Paul about my papers on function: "I haven't read his papers on function because they don't deal with the practical issues of defining junk DNA and function in the genome."
Sure, but that doesn't mean my papers can't be helpful in your context.
Here is one way in which I think my papers can be helpful.
One of the things I argue for in those papers is my view that functional biology (including molecular biology, chemical biology, cellular biology, physiology, functional morphology, and behavior biology) can be glued together as an attempt to understand how organisms are able to exist far from thermodynamic equilibrium. The key to this attempt is the notion of function as biology role, which is, roughly speaking, the role of a part or activity of an organism in maintaining that ability. Biologists analyze organisms into systems of subsystems of subsystems of … each with specific biological roles and explain the form and activity of those systems by appeal to these biological roles. The notion of biological role is not only a crucial explanatory device, it is also crucial as a way to enable a division of labour within biology while maintaining biology's coherence. The best exposition of that view is my "Biology’s Functional Perspective: Roles, Advantages and Organization" (2013).
I believe this idea about the explanatory role of function as biological role can be of help (for instance to the book you are currently writing) to put the practical issues of defining junk DNA and function in the genome into a broader context and, in that way, to clarify what is at stake in these practical issues. You might, for example, use my ideas about the functional perspective to explain why functions as biological role are so important in biology. Then raise the question how this notion is to be operationalized in the context of the study of DNA. You might want to discuss different operationalizations. The two main criteria for evaluating proposals are the validity as an operationalization of the notion of function as biological role and its usability in the relevant context.
Be aware that explaining how a certain notion of function is critical to biology is quite different from arguing that that notion of function is the one and only right way to talk of function in biology. Actually, you might learn from my 'Four notions' that the question 'what is the right notion of function?' doesn't make sense at all. This in contrast to questions like 'how is a certain notion useful for my purposes?' and 'is this proposal to define function a valid operationalization of ....'.
I have no ready made answers to your practical issues but I do think that my view on the functional perspective helps to put those questions into context and, hence, in evaluating possible answers to those issues.
lutesuite said:
ENCODE's own written statements at the time do not distinguish between "Moran function" and "ENCODE function" and, in fact, are worded so a to imply that the latter demonstrates the former
Sure. This is exactly why I said that "the mess didn't occur because ENCODE used the wrong notion of function, but because they failed to point out that their notion of function as activity is different from the notion of function as biological role used to define junk DNA."
judmarc, I don't think that Millikan is confused. Far from that! I am a great fan of her work. I think that *Graur's* notion is confused. He says that his definition is Millikan's, but it isn't. It is a strange mixture of Millikans ideas with notions explicitly rejected by Millikan. Same for Graur's notion of causal role function. He says it is Cummins' notion but it isn't and it is so in a way that takes out the key elements of Cummins approach!
Arno, I really do wish you wouldn't use "operationalization" so much.
John Harshman says,
I think it would be better if you just explain up front that "conserved" should mean "conserved relative to neutral expectation", then explain that there's expected variance to be accounted for too. And then in the examples, including hypothetical ones, you should mention the expectation.
I hate quibbling about trivial details but, since you brought it up ...
The word "conserved" implies that we know how to detect real conservation as opposed to high similarity due to chance. Thus "conserved relative to neutral expectation" is redundant unless there's also a "conserved that's not relevant to neutral expectation." In fact, there's no such thing as long as you are using the word "conserved" correctly.
What you are complaining about is not misuse of the word "conserved" but misuse of the criteria used to detect conservation. Thus, you should have been complaining about "sequence similarity relative to time of divergence and neutral expectations."
You might have called me out on not making it clear that I knew how to detect conservation. That's not the same thing as not understand the word "conservation."
I understand that you also disagree with me about some of the examples I use. That's fine. In fact it illustrates the point I'm trying to make; namely, that there's no universal definition of biological function that satisfies everyone.
The function wars have been mostly a waste of time except for proving the ENCODE definition is ridiculous and sequence conservation is a good proxy for function.
It's time to call for an armistice.
@Georgi,
I finally got around to reading the papers you referenced. The data seems to be accurate.
Some ciliates and some nulceomorphs seem to have undergone extreme selection for reduced genome size. This includes selection for the size of introns. The introns in those species have often been reduced to less than 20 bp.
This conflicts with the value of 50 bp that I've been using as the minimal size of a spliceosomal intron. We don't know how the splicing apparatus works in those species. I'm inclined to agree with Slamovits and Keeling (2009) when they suggest the splicing apparatus has also evolved since it seems to be inefficient at removing larger introns.
I've added a footnote to my post.
Nucleomorphs are really weird in general -- if you look at their histone genes, for example, they are extremely divergent (in chlorarachniophytes, in particular, they look more conventional in cryptophytes), and in ways that suggest that the typical eukaryotic transcriptional cycle exists in a highly derived state (histone marks play key role in all of those processes). There are no Pol2 CTD tails either so that part of the transcriptional cycle is gone too.
It's definitely something to study in detail directly in the future.
Sequence conservation= function may be fine for coding sequence but I'm not so sure for non-coding sequence that a biological function might not be conserved without the sequence being obviously so.
Arno Wouters says,
My interest is in clearing up the mess created by ENCODE's failure ...
That's fine but let's make sure we don't start making excuses for ENCODE's failure. That's what the Germain et al. paper does and it's why I think it's a bad paper.
Excuse are also what the ENCODE Consortium tried in their 2014 paper (Kellis et al., 2014). A good part of that paper is devoted to challenging the most important evidence for junk DNA and trying to defend their definition of function.
They claim in that paper that their goal wasn't really to provide an estimate of the fraction of the genome that is functional but merely to supply the data that will help researchers in the future.
Contrast this with the announcement on the Sanger Institute website on Sept. 5, 2012 when the ENCODE results were published ...
The ENCODE Project, today, announces that most of what was previously considered as 'junk DNA' in the human genome is actually functional. The ENCODE Project has found that 80 per cent of the human genome sequence is linked to biological function.
http://www.sanger.ac.uk/news/view/2012-09-05-google-earth-of-biomedical-research
The Sanger Centre is a major player in ENCODE.
Statements like that make it very clear that the ENCODE researchers were attacking the idea of junk DNA and presenting evidence that most of the genome is not junk but functional.
I oppose any attempt to whitewash that ridiculous attempt by pretending that ENCODE conclusions were misrepresented by the press or by pretending they were just using a different definition of function that didn't really mean "biological function."
Arno Wouters says,
The other way (taken by Graur et al. and Doolittle et al, and, apparently, advocated by you) is to start a war on the notion of function by blaming ENCODE for using the notion of ENCODE function rather than that of Moran function (or instead of Graur function or Doolittle function).
The latter response is, in my view, not constructive.
I definitely support a war on ENCODE's ridiculous attempt to prove that most of our genome has a biological function and, therefore, is not junk.
We have not won that war. ENCODE and their naive journalist supporters have convinced the vast majority of scientists that our genome is full of biological function. This is also what the general public believes.
I think this part of the war is very constructive, and necessary.
However, the other part of the Function Wars—the part where we quibble about the exact meaning of "function"— is not constructive. Indeed, it detracts from the main battle so in that sense it is counter-productive.
Arno Wouters says,
Perhaps, but to show that they were wrong we need to discuss the evidence and the conclusions that can be drawn from that evidence, rather than confused complaints about the notion of function they used. That is, we need a substantive scientific debate (hi Paul! Thanks for you support!) instead of a quasi-philosophical one.
I guess I have been clear enough by now.
I agree with you. This is a scientific debate about the evidence for and against the role of specific DNA sequences in our genome.
For example, we want to know whether a given transcription factor binding site has a biological function or is just a random sequence that happens to correspond to a binding site. Quibbling about whether to call it a biological function or a biochemical function isn't going to help.
We need to ask whether those sites can be deleted without affect the organism; whether they are conserved; whether spurious binding is consistent with what we know about biochemistry; and whether spurious binding sites can be expected according to modern evolutionary theory.
None of those questions are going to be answered by philosophers and none of them are going to be answered by trying to come up with a rigorous definition of biological function.
I didn't make my position clear in my most recent post. I did make it clear in earlier posts and I was soundly criticized for it.
I've added an UPDATE to my post so that people can more readily understand my point.
widgets101,
That would only apply to DNA that has no sequence specific activity. One example given in the article, if memory serves, is DNA spacing between genes and transcription factor binding sites. These features require a certain number of bases between the start codon and binding sites, but the sequence doesn't really matter.
For the vast majority of cases the function of a stretch of DNA is reliant on its sequence. Even RNA genes (e.g. microRNA) derive their function from specific sequences. In fact, some widely shard microRNA genes contain some of the most highly conserved sequences in their seed regions even though they are never translated into proteins.
Eric: Other structural examples might be where the secondary structure of a functional RNA is conserved but the primary sequence isn't. I don't know if current methods for predicting RNA structure are good enough to detect conservation at that level.
Laurence A. Moran: “It is wrong to arbitrarily DEFINE function in terms of selection (for sequence) because that rules out all bulk DNA hypotheses by fiat and that's not a good way to do science.”
If that is indeed the case, why don’t you critically evaluate the bulk DNA hypotheses?
If that is indeed the case, why don’t you critically evaluate the bulk DNA hypotheses?
My critical evaluation of the bulk DNA hypotheses is that none of them are worth my time and effort.
Keep calm and ask about onions.
That's fine Larry.
But one have to wonder about what if more mammal genomes will turn out to be fully or almost fully functional? Some have, surprisingly.
See my point?
On the other hand many laboratory experiments designed to prove your point about "junk DNA" theory are inconclusive or they have failed. There is more coming in almost every day.
How are you going to combat this flood of experimental scientific research Larry? What tools are going to use to counter-attack that deluge of scientific, experimental information that totally devour your beliefs?
But one have to wonder about what if more mammal genomes will turn out to be fully or almost fully functional? Some have, surprisingly.
Interesting. You wouldn't be able to provide a reference for this, would you? Y'know, in case some uncharitable individuals might think you're just making shit up.
And what is the size of the genome of this remarkable mammal whose genome has been proven to be 100% functional, compared to that of the onion? I could see some decidedly uncomfortable questions arising for you from that, regardless of the answer.
Don Quixote asks,
How are you going to combat this flood of experimental scientific research Larry? What tools are going to use to counter-attack that deluge of scientific, experimental information that totally devour your beliefs?
If, by any chance, there was experimental evidence challenging junk DNA I would use the same tools I'm using now. The tools are critical thinking, a knowledge of the scientific literature, and an understanding of fundamental concepts in biochemistry and evolution.
If, by any chance, enough genuine scientific evidence appeared to establish that most of our genome is functional, then I would accept that evidence and regret all the time I've wasted.
What tools do YOU use to distinguish truth from fiction? Evidence and critical thinking don't appear to be in your toolbox.
On the other hand many laboratory experiments designed to prove your point about "junk DNA" theory are inconclusive or they have failed. There is more coming in almost every day.
Hi DQ -
Larry's aware of the experimental data (since he happens to be writing a book on the topic, I would guess he is far more informed on said data than you are). He's quantified from time to time what the effects on our estimates of the total amount of "junk" would be if various theories/objections turned out to be correct.
Most of these effects amount to tenths of a percentage point. So perhaps ~8.8% rather than ~8.7% of our genome might be confirmed functional if one or another of these theories/objections was confirmed by good experimental evidence.
I wouldn't hold my breath waiting for the total to approach 100%. In fact there are fundamental genetic reasons why the total cannot approach 100%. (One is that we know our genes mutate at a rate of ~100+ per generation, and if they were all essential we'd die off pretty quickly as we'd all be missing essential functions due to those mutations.)
Sorry, that last bit should have been, "We know there are ~100+ mutations per generation in human DNA, and if all those mutations were in DNA essential to correct functioning and reproduction we'd die off pretty quickly as we'd all be missing essential functions due to those mutations."
Peter,
If we are talking about stem-loop structures, then it is a mixed bag. For the loop structures there is a much higher tolerance for mutations than for the stem structures which require complementary sequence. For a single base in a loop structure there are 3 other possible bases that will work in the position opposite of that base. For stem structures there is only 1 other base that will work in the opposite strand. This also applies to RNA sequence that has to bind to conserved DNA/RNA binding sites elsewhere in the cell.
A good example of this is microRNA. For example, you can look at the sequence alignment for mir-34a here:
http://people.csail.mit.edu/akiezun/microRNAviewer/all_mir-34a-align.html
The areas in blue are the seed regions that bind to messenger RNA and inhibit translation. These regions also form the stem structure in the pre-miRNA stem-loop structure. The majority of divergence occurs in the loop and open ends of the pre-miRNA.
Also, there are many algorithms out there for predicting RNA secondary structure:
https://en.wikipedia.org/wiki/List_of_RNA_structure_prediction_software
Laurence A. Moran: “My critical evaluation of the bulk DNA hypotheses is that none of them are worth my time and effort. Keep calm and ask about onions”
Isn't ironic that Ryan Gregory who came out with the ‘onion test’ started his scientific career by developing a DNA bulk hypothesis, the nucleotypic theory? Interestingly, his bulk DNA hypothesis has been highly appreciated by many scholars in the field, such as Ford Doolittle: see his recent paper “Is junk DNA bunk? A critique of ENCODE”: you can read it as it is available for free at: https://www.ncbi.nlm.nih.gov/pubmed/23479647.
I would expect that Doolittle read Gregory’s papers on the nucleotypic theory because they are collaborators (see the reference you used in your post above). So, what’s wrong with Doolittle’s evaluation, and have you ever published your critical evaluation in a scientific paper or here at Sandwalk?
To finish the discussion about intron size, here is a paper that just came out and that officially describes the 15-16bp introns in ciliates:
http://www.sciencedirect.com/science/article/pii/S0960982216315391
Post a Comment