More Recent Comments
Saturday, May 06, 2023
Friday, May 05, 2023
Thursday, May 04, 2023
90% of your genome is junk
If you are interested in learning more about junk DNA here's some links to relevant information.
Prologue Chapter 1: Introducing Genomes Chapter 2: The Evolution of Sloppy Genomes Chapter 3: Repetitive DNA and Mobile Genetic Elements Chapter 4: Why Don't Mutations Kill Us? Chaper 5: The Big Picture Chapter 6: How Many Genes? How Many Proteins? Chapter 7: Gene Families and the Birth and Death of Genes Chapter 8: Noncoding Genes and Junk RNA Chapter 9: The ENCODE Publicity Campaign Chapter 10: Turning Genes On and Off Chapter 11: Zen and the Art of Coping with a Sloppy GenomeTuesday, April 25, 2023
Happy DNA Day 2023!
It was 70 years ago today that the famous Watson and Crick paper was published in Nature along with papers by Franklin & Gosling and Wilkins, Stokes, & Wilson. Threre's a great deal of misinformation circulating about this discovery so I wrote up a brief history of the events based largely on Horace Freeland Judson's book The Eighth Day of Creation. Every biochemistry and molecular biology student must read this book or they don't qualify to be an informed scientist. However, if you are not a biochemistry student then you might enjoy my short version.
Some practising scientists might also enjoy refreshing their memories so they have an accurate view of what happened in case their students ask questions. 
Where Rosalind Franklin teaches Jim and Francis something about basic chemistry.
Where Jim and Francis discover the secret of life.
Here's the latest version of Rosalind Frankin's contribution written by Matthew Cobb and Nathaniel Comfort: What Rosalind Franklin truly contributed to the discovery of DNA's structure. If you want to know the accurate version of her history then this is a must-read. Cobb is working on a biography of Crick and Comfort is writing a biography of Watson.
Here are some other posts that might interest you on DNA Day.
- Rosalind Franklin Announces the Death of the Helix
- Sixty Years Ago Today: April 25, 1953
- Rosalind Franklin's Birthday
- Did Rosalind Franklin produce the first X-ray diffraction images of DNA?
- The Wilkins, Stokes and Wilson Nature paper (1953)
- The Franklin & Gosling Nature paper (1953)
- The Watson & Crick Nature Paper (1953)
Saturday, March 25, 2023
ChatGPT lies about junk DNA
I asked ChatGPT some questions about junk DNA and it made up a Francis Crick quotation and misrepresented the view of Susumu Ohno.
We have finally restored the Junk DNA article on Wikipedia. (It was deleted about ten years ago when Wikipedians decided that junk DNA doesn't exist.) One of the issues on Wikipedia is how to deal with misconceptions and misunderstandings while staying within the boundaries of Wikipedia culture. Wikipedians have an aversion to anything that looks like editorializing so you can't just say something like, "Nobody ever said that all non-coding DNA was junk." Instead, you have to find a credible reference to someone else who said that.
I've been trying to figure out how far the misunderstandings of junk DNA have spread so I asked ChatGPt (from OpenAI) again.
Wednesday, March 08, 2023
A small crustacean with a very big genome
The antarctic krill genome is the largest animal genome sequenced to date.
Antarctic krill (Euphausia superba) is a species of small crustacean (about 6 cm long) that lives in large swarms in the seas around Antarctica. It is one of the most abundant animals on the planet in terms of biomass and numbers of individuals.
It was known to have a large genome with abundant repetitive DNA sequences making assembly of a complete genome very difficult. Recent technological advances have made it possible to sequence very long fragments of DNA that span many of the repetitive regions and allow assembly of a complete genome (Shao et al. 2023).
The project involved 28 scientists from China (mostly), Australia, Denmark, and Italy. To give you an idea of the effort involved, they listed the sequencing data that was collected: 3.06 terabases (Tb) PacBio long read sequences, 734.99 Gb PacBio circular consensus sequences, 4.01 Tb short reads, and 11.38 Tb Hi-C reads. The assembled genome is 48.1 Gb, which is considerably larger than that of the African lungfish (40 Gb), which up until now was the largest fully sequenced animal genome.
The current draft has 28,834 protein-coding genes and an unknown number of noncoding genes. About 92% of the genome is repetitive DNA that's mostly transposon-related sequences. However, there is an unusual amount of highly repetitive DNA organized as long tandem repeats and this made the assembly of the complete genome quite challenging.
The protein-coding genes in the Antarctic krill are longer than in other species due to the insertion of repetitive DNA into introns but the increase in intron size is less than expected from studies of other large genomes such as lungfish and Mexican axolotl. It looks like more of the genome expansion has occurred in the intergenic DNA compared to these other species.
This study supports the idea that genome expansion is mostly due to the insertion and propagation of repetitive DNA sequences. Some of us think that the repetitive DNA is mostly junk DNA but in this case it seems unusual that there would be so much junk in the genome of a species with such a huge population size (about 350 trillion individuals). The authors were aware of this problem but they were able to calculate an effective population size because they had sequence data from different individuals all around Antarctica. The effective population size (Ne) turned out to be one billion times smaller than the census population size indicating that the population of krill had been much smaller in the recent past. Their data suggests strongly that this smaller population existed only 10 million years ago.
The authors don't mention junk DNA. They seem to favor the idea that large genomes are associated with crustaceans that live in polar regions and that large genomes may confer a selective advantage.
Shao, C., Sun, S., Liu, K., Wang, J., Li, S., Liu, Q., Deagle, B.E., Seim, I., Biscontin, A., Wang, Q. et al. (2023) The enormous repetitive Antarctic krill genome reveals environmental adaptations and population insights. Cell 186:1-16. [doi: 10.1016/j.cell.2023.02.005]
Friday, March 03, 2023
Do you understand the scientific literature?
I'm finding it increasingly difficult to understand the scientific literature even in subjects that I've been following for decades. Is it just because I'm getting too old to keep up?
Here's an example of a paper that I'd like to understand but after reading the abstract and the introduction I gave up. I'll quote the first paragraph of the introduction to see if any Sandwalk readers can do better.
I'm not talking about the paper being a complete mystery; I can figure out roughly what's it's about. What I'm thinking is that the opening paragraph could have been written in a way that makes the goals of the research much more comprehensible to average scientifically-literate people.
Weiner, D. J., Nadig, A., Jagadeesh, K. A., Dey, K. K., Neale, B. M., Robinson, E. B., ... & O’Connor, L. J. (2023) Polygenic architecture of rare coding variation across 394,783 exomes. Nature 614:492-499. [doi = 10.1038/s41586-022-05684-z]
Genome-wide association studies (GWAS) have identified thousands of common variants that are associated with common diseases and traits. Common variants have small effect sizes individually, but they combine to explain a large fraction of common disease heritability. More recently, sequencing studies have identified hundreds of genes containing rare coding variants, and these variants can have much larger effect sizes. However, it is unclear how much heritability rare variants explain in aggregate, or more generally, how common-variant and rare-variant architecture compare: whether they are equally polygenic; whether they implicate the same genes, cell types and genetically correlated risk factors; and whether rare variants will contribute meaningfully to population risk stratification.
The first question that comes to mind is whether the variant that's associated with a common disease is the cause of that disease or merely linked to the actual cause. In other words, are the associated variants responsible for the "effect size"? It sounds like the answer is "yes" in this case. Has that been firmly esablished in the GWAS field?
Thursday, March 02, 2023
"You like me!"
The endorsements for my book are in.
One of the last steps in publishing a book is to collect endorsements—favorable statements from famous people who urge you to buy the book. These short endorsements will appear in the front of the book and on the book jacket (dust jacket). They may also appear on various websites in order to promote sales.
The trick is to sent the book out for review to as many people as possible and hope that one or two will like it well enough to say something nice. I'm pleased to report that there were, indeed, a few people who liked the book well enough to endorse it.
The title of this post is from Sally Field's acceptance speech on winning the Academy Award for best actress in 1985. She said, "I can't deny the fact that you like me. Right now, you like me!"
Wednesday, March 01, 2023
Definition of a gene (again)
The correct definition of a molecular gene isn't difficult but getting it recognized and accepted is a different story.
When writing my book on junk DNA I realized that there was an issue with genes. The average scientist, and consequently the average science writer, has a very confused picture of genes and the proper way to define them. The issue shouldn't be confusing for Sandwalk readers since we've covered that ground many times in the past. I think the best working definition of a gene is, "A gene is a DNA sequence that is transcribed to produce a functional product" [What Is a Gene?]
Saturday, February 25, 2023
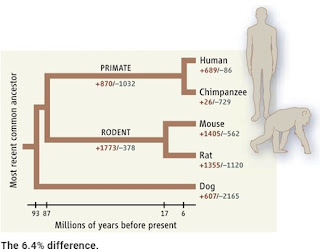
How Intelligent Design Creationists try to deal with the similarity between human and chimp genomes
The initial measurement of the difference between the human and chimp genomes was based on aligning 2.4 billion base pairs in the two genomes. This gave a difference of 1.23% by counting base pair substitutions and small deletions and insertions (indels). However, if you look at larger indels, including genes, you can come up with bigger values because you can count the total number of base pairs in each indel; for example, a deletion of 1,000 bp will be equivalent to 1,000 SNPs.
Thursday, February 16, 2023
What are the best Nobel Prizes in biochemistry & molecular biology since 1945?
The 2022 Nobel Prize in Physiology or Medicne went to Svante Pääbo “for his discoveries concerning the genomes of extinct hominins and human evolution”. It's one of a long list of Nobel Prizes awarded for technological achievement. It most cases, the new techniques led to a better understanding of science and medicine.
Since World War II, there have been significant advances in our understanding of biology but most of these have come about by the slow and steady accumulation of knowledge and not by paradigm-shifting breakthroughs. These advances don't often get recognized by the Nobel Prize committees because it's difficult to single out any one individual or any single experiment that merits a Nobel Prize. In some cases the Nobel Prize committees have tried to recognize major advances by picking out leaders that have made important contributions over a number of years but their choices don't always satisfy others in the field. One of the notable successes is the awarding of Nobel Prizes to Max Delbrück, Alfred D. Hershey and Salvador E. Luria “for their discoveries concerning the replication mechanism and the genetic structure of viruses” (Nobel Prize in Physiology or Medicine 1969). Another is Edward B. Lewis, Christiane Nüsslein-Volhard and Eric F. Wieschaus “for their discoveries concerning the genetic control of early embryonic development” (Nobel Prize in Physiology or Medicine 1995)
Birds of a feather: epigenetics and opposition to junk DNA
There's an old saying that birds of a feather flock together. It means that people with the same interests tend to associate with each other. It's extended meaning refers to the fact that people who believe in one thing (X) tend to also believe in another (Y). It usually means that X and Y are both questionable beliefs and it's not clear why they should be associated.
I've noticed an association between those who promote epigenetics far beyond it's reasonable limits and those who reject junk DNA in favor of a genome that's mostly functional. There's no obvious reason why these two beliefs should be associated with each other but they are. I assume it's related to the idea that both beliefs are presumed to be radical departures from the standard dogma so they reinforce the idea that the author is a revolutionary.
Or maybe it's just that sloppy thinking in one field means that sloppy thinking is the common thread.
Here's an example from Chapter 4 of a 2023 edition of the Handbook of Epigenetics (Third Edition).
The central dogma of life had clearly established the importance of the RNA molecule in the flow of genetic information. The understanding of transcription and translation processes further elucidated three distinct classes of RNA: mRNA, tRNA and rRNA. mRNA carries the information from DNA and gets translated to structural or functional proteins; hence, they are referred to as the coding RNA (RNA which codes for proteins). tRNA and rRNA help in the process of translation among other functions. A major part of the DNA, however, does not code for proteins and was previously referred to as junk DNA. The scientists started realizing the role of the junk DNA in the late 1990s and the ENCODE project, initiated in 2003, proved the significance of junk DNA beyond any doubt. Many RNA types are now known to be transcribed from DNA in the same way as mRNA, but unlike mRNA they do not get translated into any protein; hence, they are collectively referred to as noncoding RNA (ncRNA). The studies have revealed that up to 90% of the eukaryotic genome is transcribed but only 1%–2% of these transcripts code for proteins, the rest all are ncRNAs. The ncRNAs less than 200 nucleotides are called small noncoding RNAs and greater than 200 nucleotides are called long noncoding RNAs (lncRNAs).
In case you haven't been following my blog posts for the past 17 years, allow me to briefly summarize the flaws in that paragraph.
- The central dogma has nothing to do with whether most of our genome is junk
- There was never, ever, a time when knowledgeable scientists defended the idea that all noncoding DNA is junk
- ENCODE did not "prove the significance of junk DNA beyond any doubt"
- Not all transcripts are functional; most of them are junk RNA transcribed from junk DNA
So, I ask the same question that I've been asking for decades. How does this stuff get published?
Wednesday, February 15, 2023
The Maud Menten Center
Several people have commented on Facebook about an interview with Matthew Cobb [Reflections on the Double Helix's Platinum Anniversary]. Cobb has written several books that you all should have read by now. You may also be familiar with his name from Jerry Coyne's website since he is mentioned there quite a lot.
Cobb is currently writing a biography of Francis Crick and it's the reference to Crick that's attracting attention.
Watson and Crick had something that very few of us have, which is a very relaxed relationship that allowed arguing and talking and debating and being prepared to have crazy ideas that the other person could shoot down.
This is something that Crick continued to do with Nobelist Sydney Brenner, PhD, for over 20 years. Every morning, they would just yak at each other for hours on end and talk rubbish. And most of it was rubbish! But every now and again, there would be something really insightful that they had seen in an article that they could develop further.
And that’s not exactly possible in today’s world, which focuses on how many Nature, Science, or Cell articles are on a CV. So, they were living in a very different world that was much more relaxed. But I think if we could reinject a bit of that back into science—more freedom and time to explore—I think everybody would benefit.
I think we need an institute that's similar to the Institute for Advanced Studies in Princeton except that it would be focused on biochemistry and molecular biology, including genomics and molecular evolution. It would be a place for theorists and it would encourage the kind of interactions that Cobb is talking about. I think we should create that institute in Toronto and call it the Maud Menten Center after Maud L. Menten who got her undergraduate medical degree at the University of Toronto in 1907 and later on (1911) was awarded the advanced medical degree equivalent to a Ph.D.1 Most of you will know her from her work on the theory of enzyme kinetics with Leonor Michaelis. Michaelis-Menten kinetics is one of the most important contributions to biochemistry in the 20th century.
The Maud Menten Center should have a permanent staff of several smart scientists and facilities for a large number of visiting scientists who could spend their sabbaticals at the institute. There are major corporations in Toronto that could be encouraged to contribute to supporting research in this field but a lot of the support might come from various levels of government. The Canadian model is the Perimeter Institute for Theoretical Physics in Waterloo, Ontario, Canada.
1. There's a lot of confusion about the various degrees Menten got at the University of Toronto but with the help of a few people in the alumni office we were able to sort it out [The mystery of Maud Menten].
David Allis (1951 - 2023) and the "histone code"
C. David Allis died on January 8, 2023. You can read about his history of awards and accomplishments in the Nature obituary with the provocative subtitle Biologist who revolutionized the chromatin and gene-expression field. This refers to his work on histone acetyltransferases (HATs) and his ideas about the histone code.
The key paper on the histone code is,
Strahl, B. D., and Allis, C. D. (2000) The language of covalent histone modifications. Nature, 403:41-45. [doi: 10.1038/47412]
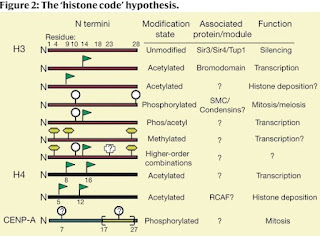
Histone proteins and the nucleosomes they form with DNA are the fundamental building blocks of eukaryotic chromatin. A diverse array of post-translational modifications that often occur on tail domains of these proteins has been well documented. Although the function of these highly conserved modifications has remained elusive, converging biochemical and genetic evidence suggests functions in several chromatin-based processes. We propose that distinct histone modifications, on one or more tails, act sequentially or in combination to form a ‘histone code’ that is, read by other proteins to bring about distinct downstream events.
They are proposing that the various modifications of histone proteins can be read as a sort of code that's recognized by other factors that bind to nucleosomes and regulation gene expression.
This is an important contribution to our understanding of the relationship between chromatin structure and gene expression. Nobody doubts that transcription is associated with an open form of chromatin that correlates with demethylation of DNA and covalent modifications of histone and nobody doubts that there are proteins that recognize modified histones. However, the key question is what comes first; the binding of transcription factors followed by changes to the DNA and histones, or do the changes to DNA and histones open the chromatin so that transcription factors can bind? These two models are referred to as the histone code model and the recruitment model.
Strahl and Allis did not address this controversy in their original paper; instead, they concentrated on what happens after histones become modified. That's what they mean by "downstream events." Unfortunately, the histone code model has been appropriated by the epigenetics cult and they do not distinguish between cause and effect. For example,
The “histone code” is a hypothesis which states that DNA transcription is largely regulated by post-translational modifications to these histone proteins. Through these mechanisms, a person’s phenotype can change without changing their underlying genetic makeup, controlling gene expression. (Shahid et al. (2022)
The language used by fans of epigenetics strongly implies that it's the modification of DNA and histones that is the primary event in regulating gene expression and not the sequence of DNA. The recruitment model states that regulation is primarily due to the binding of transcription factors to specific DNA sequences that control regulation and then lead to the epiphenomenon of DNA and histone modification.
The unauthorized expropriation of the histone code hypothesis should not be allowed to diminish the contribution of David Allis.
Wikipedia vs experts and a proposal for "arbitrators"
Wikipedia is a not-for-profit crowdsourced encyclopedia that's open to anybody who wants to contribute. This is both a strength and a weakness but the weaknesses are becoming important in an age of fake news and misinformation. The rules of Wikipedia mean that amateurs can insert any information into science articles as long as it is backed by a reliable source. But "reliable sources" include the popular press and books that may or may not report the scientific consensus accurately. When knowledgeable experts try to correct information, or put it into the proper context, they are often opposed by Wikipedia administrators who have a built-in bias against experts—a bias that's not entirely unjustified but much abused. Consequently, scientists often get frustrated trying to deal with the rules and traditions of Wikipedians because these rules are very different than the standards in the scientific community.
Here's an interesting article by Piotr Konieczny on From Adversaries to Allies? The Uneasy Relationship between Experts and the Wikipedia Community.