New techniques are being developed to obtain the complete sequences of both copies (maternal and paternal) of a typical diploid individual.
The first two sequences of the human genome were published twenty years ago by the International Human Genome Project and by a company called Celera Genomics. The published sequences were a consensus using DNA from multiple indivduals so the final result didn't represent the sequence of any one person. Furthermore, since each of us has inherited separate genomes from our mother and father, our DNA is actually a mixture of two different haploid genomes. Most published genome sequences are an average of these two separate genomes where the choice of nucleotide at any one position is arbitrary.
The first person to have a complete genome sequence was James Watson in 2007 but that was a composite genome sequence. Craig Venter's genome sequence was published a few months later and it was the first complete genome sequence containing separate sequences of each of his 46 chromosomes. (One chromosome from each of his parents.) In today's language, we refer to this as a diploid sequence.
The current reference sequence is based on the data published by the public consortium (International Humand Genome Project)—nobody cares about the Celera sequence. Over the years, more and more sequencing data has been published and this has been incorporated into the standard human reference genome in order to close most gaps and improve the accuracy. The current version is called GRCh38.p14 from February 3, 2022. It's only 95% complete because it's missing large stretches of repetitive DNA, especially in the centromere regions and at the ends of each chromosome (telomeric region).
The important point for this discussion is that CRCh38 is not representative of the genomes of most people on Earth because there has been a bias in favor of sequencing European genomes. (Some variants are annotated in the reference genome but this can't continue.) Many scientists are interested the different kinds of variants present in the human population so they would like to create databases of genomes from diverse populations.
The first complete, telomere-to-telomere (T2T), human genome sequence was published last year [A complete human genome sequence (2022). It was made possible by advances in sequencing technology that generated long reads of 10,000 bp and ultra-long reads of up to 1,000,000 bp [Telomere-to-telomere sequencing of a complete human genome]. The DNA is from a CHM13 cell line that has identical copies of each chromosome so there's no ambiguity due to differences in the maternal and paternal copies. The full name of this sequence is CHM13-T2T.
The two genomes (CRCh38 and CHM13) can't be easily merged so right now there are competing reference genomes [What do we do with two different human genome reference sequences?].
The techniques used to sequence the CHM13 genome make it possible to routinely obtain diploid genome sequences from a large number of individuals because overlapping long reads can link markers on the same chromosome and distinguish between the maternal and paternal chromosomes. However, in practice, the error rate of long read sequencing made assembly of separate chromosomes quite difficult. Recent advances in the accuracy of long read sequencing have been developed by PacBio, and this high fidelity sequencing (PacBio HiFi sequencing) promises to change the game.
The Human Pangene Reference Consortium has tackled the problem by sequencing the genome of an Ashkenazi man (HG002) and his parents (HG002-father and HG004-mother) using the latest sequencing techniques. They then asked the genome community to submit their assemblies using their best software in a kind of "assembly bakeoff." They got 23 responses.
Jarvis, E. D., Formenti, G., Rhie, A., Guarracino, A., Yang, C., Wood, J., et al. (2022) Semi-automated assembly of high-quality diploid human reference genomes. Nature, 611:519-531. [doi: 10.1038/s41586-022-05325-5]
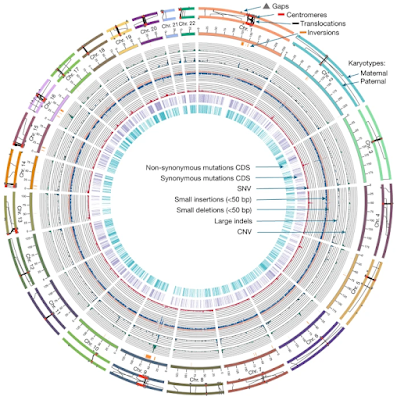
The current human reference genome, GRCh38, represents over 20 years of effort to generate a high-quality assembly, which has benefitted society. However, it still has many gaps and errors, and does not represent a biological genome as it is a blend of multiple individuals. Recently, a high-quality telomere-to-telomere reference, CHM13, was generated with the latest long-read technologies, but it was derived from a hydatidiform mole cell line with a nearly homozygous genome. To address these limitations, the Human Pangenome Reference Consortium formed with the goal of creating high-quality, cost-effective, diploid genome assemblies for a pangenome reference that represents human genetic diversity. Here, in our first scientific report, we determined which combination of current genome sequencing and assembly approaches yield the most complete and accurate diploid genome assembly with minimal manual curation. Approaches that used highly accurate long reads and parent–child data with graph-based haplotype phasing during assembly outperformed those that did not. Developing a combination of the top-performing methods, we generated our first high-quality diploid reference assembly, containing only approximately four gaps per chromosome on average, with most chromosomes within ±1% of the length of CHM13. Nearly 48% of protein-coding genes have non-synonymous amino acid changes between haplotypes, and centromeric regions showed the highest diversity. Our findings serve as a foundation for assembling near-complete diploid human genomes at scale for a pangenome reference to capture global genetic variation from single nucleotides to structural rearrangements.
We don't need to get into all the details but there are a few observations of interest.
- All of the attempted assemblies were reasonably good but the best ones had to make use of the parental genomes to resolve discrepancies.
- Some assemblies began by separating the HG002 (child) sequences into two separate groups based on their similarity to one of the parents. Others generated assemblies without using the parental data then fixed any problems by using the parental genomes and a technique called "graph-based phasing." The second approach was better.
- All of the final assemblies were contaminated with varying amounts of E. coli and yeast DNA or and/or various adaptor DNA sequences that were not removed by filters. All of them were contaminated with mitochondrial DNA that did not belong in the assembled chromosomes.
- The most common sources of assembly errors were: (1) missing joins where large stretches of DNA should have been brought together, (2) misjoins where two large stretches (contigs) were inappropriately joined, (3) incorrect inversions, and (4) false duplications.
- The overall accuracy of the best assemblies was one base pair error in 100,000 bp (10-5).
- Using the RefSeq database of 27,225 genes, most assemblies captured almost all of these confirmed and probable genes but several hundred were not complete and many were missing.
- No chromosome was complete telomere-telomere (T2T) but most were nearly complete including the complicated centromere and telomere regions.
- The two genomes (parental and maternal) differed at 2.6 million SNPs (single nucleotides), 631,000 small structural variations (<50 bp), and 11,600 large structural variations (>50 bp).
- The consortium used the best assembly algorithm to analyze the genomes of an additional 47 individuals. They began with the same coverage used for HG002; namely, 35X coverage. (Each stretch of DNA was sequenced 35 times on average - about equal amounts in both directions.) This was not successful so they had to increase the coverage to 130X to get good assemblies. They estimate that each additional diploid sequence will reguire 50-60X coverage. This kind of coverage would have been impossible in the 1990s when the first human genome was assembled but now it's fairly easy as long as you have the computer power and storage to deal with it.

Thanks for this review.
ReplyDelete"The overall accuracy of the best assemblies was one base pair error in 100,000 bp"
ReplyDeletethat would imply that the best assemblies contain 35.000 errors per haploid genome?
@gert korthif
ReplyDeleteThat's correct. The average assembly has about 30,000 errors per haploid genome. That's a typical error rate for most genomes that have been carefully done but the literature is full of genome sequences that are of the "quick and dirty" type with much higher error rates.