A recent paper in Nature claims that 75% of synonymous mutations reduce fitness in yeast. The results were challenged (refuted?) ten weeks later in a manuscript posted on a preprint server.
The first paper was published in the June 23, 2022 issue of Nature (Shen et al., 2022). The authors looked at mutations in 21 nonessential genes in yeast where mutations are known to lower fitness. They created mutations in the coding regions of these genes using the CRISPR-Cas9 editing technique. A total of 1,866 synonymous mutations were created as well as 6,306 non-synonymous mutations and 169 nonsense mutations.
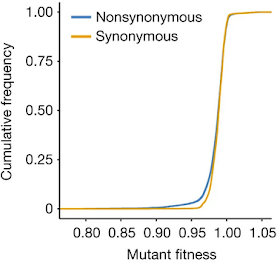
Each mutant strain was grown in rich medium at 30° C with a wild-type strain and the fitness of the mutant strain was judged by how well it competed against the wild-type strain. The mean fitness level of the nonsense mutation strains was 0.940 and the average fitness of the non-synonymous strains was 0.988 indicating that they are more fit than the strains with the nonsense mutations.
Surprisingly, the mean fitness level of the synonymous mutation strains was 0.989, which indicates that the average synonymous mutation reduces fitness as much as a nonsynonymous mutation. Closer examination of individual mutations revealed that 75.9% of synonymous mutations are "significantly deleterious" and 1.3% are "significantly beneficial." The corresponding values for nonsynonymous mutations are 75.8% and 1.6% respectively.
Some experiments suggest that the deleterious effect of synonymous mutations might be due to decreased mRNA stability but this doesn't account for the lowered fitness of all of the synonymous mutations.
The take-home message is that most synonymous mutations are non-neutral and this result conflicts with hundreds of studies done over the past 50 years that, collectively, have shown that most (but not all) synonymous mutations are neutral or nearly neutral. In one of the greatest understatements in recent memory, the authors say,
Because many biological conclusions rely on the presumption that synonymous mutations are neutral or nearly neutral, its invalidation has broad implications.
I was going to write a post about this paper last June but it didn't look right to me. The differences in fitness seemed insignificant and I was worried about whether there was something unusual about the genes they selected.
Kruglyak et al. (2022) have posted a preprint on the same topic. Their title gives a hint concerning how they feel about the Nature paper: "No evidence that synonymous mutations in yeast genes are mostly deleterious."
These authors (Kruglyak et al.) bring up a significant point when they say,
... we note that when results appear to contradict decades of evidence from multiple fields, the default assumption should be that they arise from a hidden artifact or artifacts, and every effort should be made to rule these out before taking the results at face value.
The essence of their objection to the Shen et al. paper can be summarized in four points.
- The mutant strains may have additional defects caused by transformations.
- The mutant strains may have additional defects caused by deleting certain genes during preparation of the strains. "The combination of all of these effects means that strains in which each of the 21 genes were first deleted may have fitness differences from each other and from the single WT control strain, making it impossible to properly assign fitness to the individual engineered mutations.
- The CRISPR-Cas9 insertions may have created invisible fitness-reducing mutations.
- The minor differences in fitness differences and some anomalies in the data suggest a "potential bias in one of the assays."
But their main objection is the selection of the so-called wild-type strain. They don't believe that it serves as a valid control for the mutant strains since it was picked as a cloned, individual, healthy grower from a heterogeneous population. Thus Shen et al. may have inadvertently selected a "control" strain that grows a bit better than any of the original parental strains that were mutated.
The selection of a single WT strain is especially sensitive to potential ascertainment bias, as any obviously unhealthy strain would not be selected to represent WT fitness, but within the variant pools and among pools of the same variant there is no such filter.
The lead author on this paper joined with several other scientists who signed a letter that appears to be intended for publication in Nature [A minimal role for synonymous variation in human disease]. They say,
"Synonymous mutations change the DNA sequence of a gene without affecting the amino acid sequence of the encoded protein. Although emerging evidence suggests that synonymous mutations can impact RNA splicing, translational efficiency, and mRNA stability, studies in human genetics, mutagenesis screens, and other experiments and evolutionary analyses have repeatedly shown that most synonymous variants are neutral or only weakly deleterious, with some notable exceptions. In their recent article, Shen et al. claim to have disproved these well-established findings. They perform mutagenesis experiments in yeast and conclude that synonymous mutations frequently reduce fitness to the same extent as nonsynonymous mutations. Based on their findings, the authors state that their results “imply that synonymous mutations are nearly as important as nonsynonymous mutations in causing disease.” An accompanying News and Views argues that “revising our expectations about synonymous mutations should expand our view of the genetic underpinnings of human health”. Considering potential technical concerns with these experiments and a large, coherent body of knowledge establishing the predominant neutrality of synonymous variants, we caution against interpreting this study in the context of human disease."
The arguments of Kruglyak are more developed in the manuscript than I'm able to summarize here. They seem reasonable to me, suggesting that the reviewers of the Nature paper failed to exercise the proper scrutiny of a paper that contradicted many previous studies. I suspect that this reflects a desire among Nature editors to publish "breakthrough" papers and that this may create an environment where healthy skepticism is downplayed.
Or, maybe Nature just has bad editors and bad reviewers.
Kruglyak, L., Beyer, A., Bloom, J. S., Grossbach, J., Lieberman, T. D., Mancuso, C. P., Rich, M.S., Sherlock, G., van Nimwegen, E. and Kaplan, C. D. (2022) No evidence that synonymous mutations in yeast genes are mostly deleterious. bioRxiv. [doi: 10.1101/2022.07.14.500130]
Shen, X., Song, S., Li, C., and Zhang, J. (2022) Synonymous mutations in representative yeast genes are mostly strongly non-neutral. Nature 606:725-731. [doi: 10.1038/s41586-022-04823-w]

Another challenge to the same paper (or more precisely, its accompanying News and Views) from the perspective of human disease genetics:
ReplyDelete"A minimal role for synonymous variation in human disease"
https://www.biorxiv.org/content/10.1101/2022.07.13.499964v1
Good luck getting Nature to retract. I had a run in with them several years ago over a front page story. We pointed out that the structures could better be explained by everyday processes. The editors rejected our correspondence because it contained "nothing new" even though one of the reviewers admitted that the structures were what we said they were. Presumably only new science can refute a Nature paper. Chris Nedin (University of Ediacara)
ReplyDelete@Anonymous #1
ReplyDeleteThanks for the link. The Kruglyak et al. paper mentioned the letter but I couldn't find it.
I put it in my blog post.
For an update by Shen et al., please see: bioRxiv preprint doi: https://doi.org/10.1101/2022.08.22.504687; this version posted August 23, 2022.
ReplyDelete